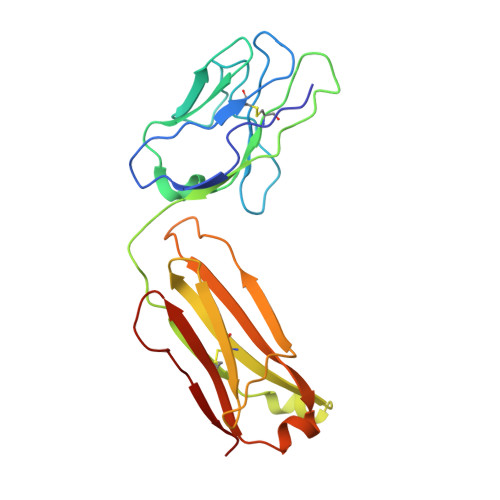
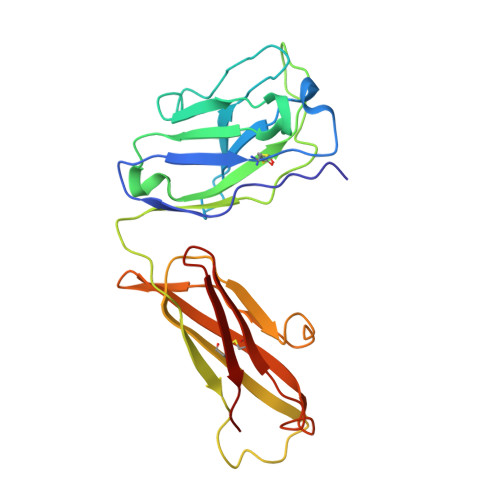
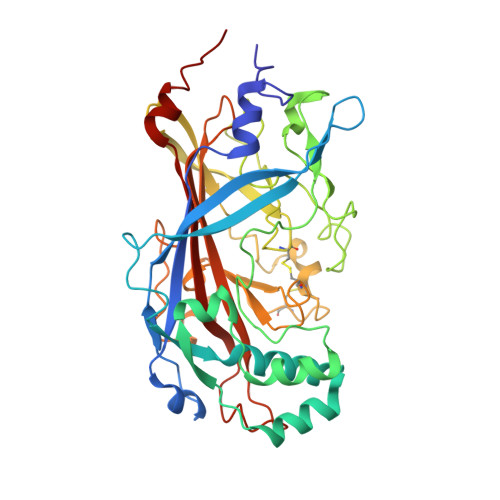
The binding of a monoclonal antibody to the apical region of SCARB2 blocks EV71 infection.
Zhang, X., Yang, P., Wang, N., Zhang, J., Li, J., Guo, H., Yin, X., Rao, Z., Wang, X., Zhang, L.(2017) Protein Cell 8: 590-600
- PubMed: 28447294
- DOI: https://doi.org/10.1007/s13238-017-0405-7
- Primary Citation of Related Structures:
5XBM - PubMed Abstract:
Entero virus 71 (EV71) causes hand, foot, and mouth disease (HFMD) and occasionally leads to severe neurological complications and even death. Scavenger receptor class B member 2 (SCARB2) is a functional receptor for EV71, that mediates viral attachment, internalization, and uncoating. However, the exact binding site of EV71 on SCARB2 is unknown. In this study, we generated a monoclonal antibody (mAb) that binds to human but not mouse SCARB2. It is named JL2, and it can effectively inhibit EV71 infection of target cells. Using a set of chimeras of human and mouse SCARB2, we identified that the region containing residues 77-113 of human SCARB2 contributes significantly to JL2 binding. The structure of the SCARB2-JL2 complex revealed that JL2 binds to the apical region of SCARB2 involving α-helices 2, 5, and 14. Our results provide new insights into the potential binding sites for EV71 on SCARB2 and the molecular mechanism of EV71 entry.
Organizational Affiliation:
Key Laboratory of Infection and Immunity, Institute of Biophysics, Chinese Academy of Sciences, Beijing, 100101, China.