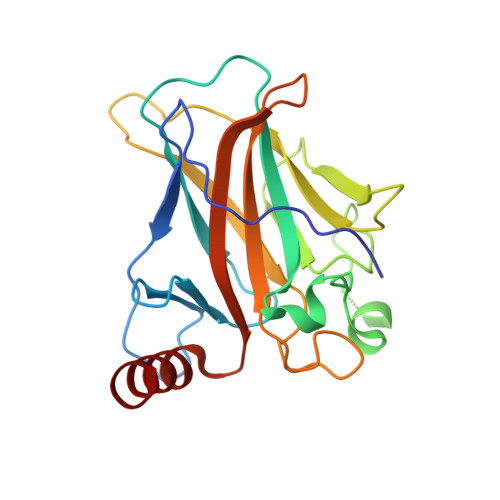
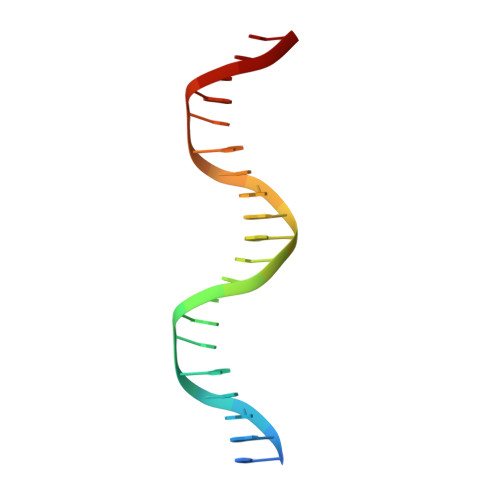
New Insights into the Role of DNA Shape on Its Recognition by p53 Proteins.
Golovenko, D., Brauning, B., Vyas, P., Haran, T.E., Rozenberg, H., Shakked, Z.(2018) Structure 26: 1237-1250.e6
- PubMed: 30057026
- DOI: https://doi.org/10.1016/j.str.2018.06.006
- Primary Citation of Related Structures:
5MCT, 5MCU, 5MCV, 5MCW, 5MF7, 5MG7, 6FJ5 - PubMed Abstract:
The tumor suppressor p53 acts as a transcription factor recognizing diverse DNA response elements (REs). Previous structural studies of p53-DNA complexes revealed non-canonical Hoogsteen geometry of A/T base pairs at conserved CATG motifs leading to changes in DNA shape and its interface with p53. To study the effects of DNA shape on binding characteristics, we designed REs with modified base pairs "locked" into either Hoogsteen or Watson-Crick form. Here we present crystal structures of these complexes and their thermodynamic and kinetic parameters, demonstrating that complexes with Hoogsteen base pairs are stabilized relative to those with all-Watson-Crick base pairs. CATG motifs are abundant in p53REs such as GADD45 and p53R2 related to cell-cycle arrest and DNA repair. The high-resolution structures of these complexes validate their propensity to adopt the unique Hoogsteen-induced structure, thus providing insights into the functional role of DNA shape and broadening the mechanisms that contribute to DNA recognition by proteins.
Organizational Affiliation:
Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.