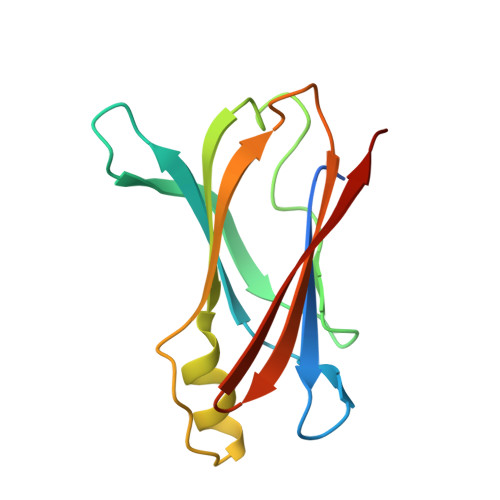
Cavity filling mutations at the thyroxine-binding site dramatically increase transthyretin stability and prevent its aggregation.
Sant'Anna, R., Almeida, M.R., Varejao, N., Gallego, P., Esperante, S., Ferreira, P., Pereira-Henriques, A., Palhano, F.L., de Carvalho, M., Foguel, D., Reverter, D., Saraiva, M.J., Ventura, S.(2017) Sci Rep 7: 44709-44709
- PubMed: 28338000
- DOI: https://doi.org/10.1038/srep44709
- Primary Citation of Related Structures:
5FO2, 5FW6 - PubMed Abstract:
More than a hundred different Transthyretin (TTR) mutations are associated with fatal systemic amyloidoses. They destabilize the protein tetrameric structure and promote the extracellular deposition of TTR as pathological amyloid fibrils. So far, only mutations R104H and T119M have been shown to stabilize significantly TTR, acting as disease suppressors. We describe a novel A108V non-pathogenic mutation found in a Portuguese subject. This variant is more stable than wild type TTR both in vitro and in human plasma, a feature that prevents its aggregation. The crystal structure of A108V reveals that this stabilization comes from novel intra and inter subunit contacts involving the thyroxine (T 4 ) binding site. Exploiting this observation, we engineered a A108I mutation that fills the T 4 binding cavity, as evidenced in the crystal structure. This synthetic protein becomes one of the most stable TTR variants described so far, with potential application in gene and protein replacement therapies.
Organizational Affiliation:
Institut de Biotecnologia i Biomedicina and Departament de Bioquímica i Biologia Molecular, Universitat Autònoma de Barcelona, Bellaterra, Spain.