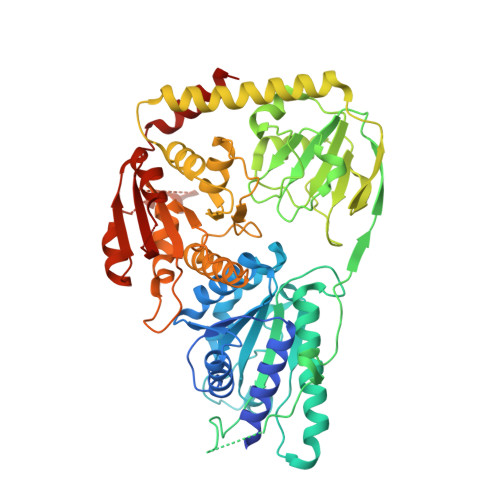
Structural Views along the Mycobacterium tuberculosis MenD Reaction Pathway Illuminate Key Aspects of Thiamin Diphosphate-Dependent Enzyme Mechanisms.
Jirgis, E.N., Bashiri, G., Bulloch, E.M., Johnston, J.M., Baker, E.N.(2016) Structure 24: 1167-1177
- PubMed: 27291649
- DOI: https://doi.org/10.1016/j.str.2016.04.018
- Primary Citation of Related Structures:
5ERX, 5ERY, 5ESD, 5ESO, 5ESS, 5ESU - PubMed Abstract:
Menaquinone (MQ) is an essential component of the respiratory chains of many pathogenic organisms, including Mycobacterium tuberculosis (Mtb). The first committed step in MQ biosynthesis is catalyzed by 2-succinyl-5-enolpyruvyl-6-hydroxy-3-cyclohexadiene-1-carboxylate synthase (MenD), a thiamin diphosphate (ThDP)-dependent enzyme. Catalysis proceeds through two covalent intermediates as the substrates 2-oxoglutarate and isochorismate are successively added to the cofactor before final cleavage of the product. We have determined a series of crystal structures of Mtb-MenD that map the binding of both substrates, visualizing each step in the MenD catalytic cycle, including both intermediates. ThDP binding induces a marked asymmetry between the coupled active sites of each dimer, and possible mechanisms of communication can be identified. The crystal structures also reveal conformational features of the two intermediates that facilitate reaction but prevent premature product release. These data fully map chemical space to inform early-stage drug discovery targeting MenD.
Organizational Affiliation:
Maurice Wilkins Centre for Molecular Biodiscovery, School of Biological Sciences, University of Auckland, Private Bag 92019, Auckland 1010, New Zealand.