Optimization of a Novel Binding Motif to (E)-3-(3,5-Difluoro-4-((1R,3R)-2-(2-Fluoro-2-Methylpropyl)-3-Methyl-2, 3,4,9-Tetrahydro-1H-Pyrido[3,4-B]Indol-1-Yl)Phenyl)Acrylic Acid (Azd9496), a Potent and Orally Bioavailable Selective Estrogen Receptor Downregulator and Antagonist.
De Savi, C., Bradbury, R.H., Rabow, A.A., Norman, R.A., De Almeida, C., Andrews, D.M., Ballard, P., Buttar, D., Callis, R., Currie, G.S., Davies, C., Donald, C., Feron, L., Hayter, B.R., Hussain, S., Karoutchi, G., Lamont, S., Macfaul, P.A., Moss, T.A., Pearson, S., Tonge, M., Walker, G., Weir, H., Wilson, Z.(2015) J Med Chem 58: 8128
- PubMed: 26407012
- DOI: https://doi.org/10.1021/acs.jmedchem.5b00984
- Primary Citation of Related Structures:
5AAU, 5AAV, 5ACC - PubMed Abstract:
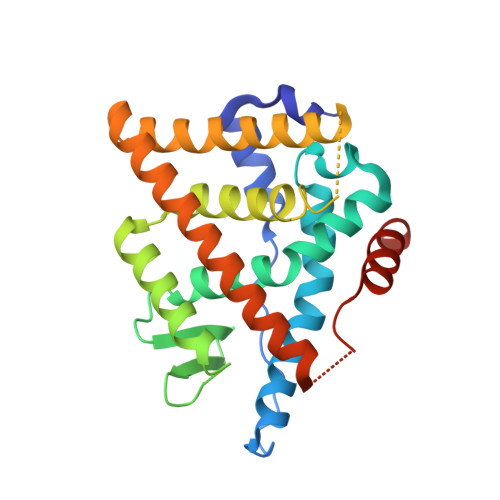
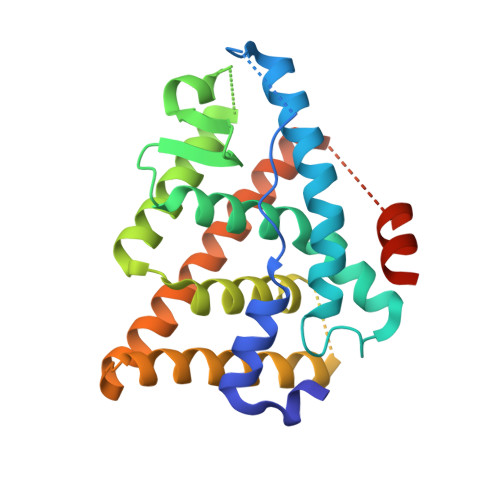
The discovery of an orally bioavailable selective estrogen receptor downregulator (SERD) with equivalent potency and preclinical pharmacology to the intramuscular SERD fulvestrant is described. A directed screen identified the 1-aryl-2,3,4,9-tetrahydro-1H-pyrido[3,4-b]indole motif as a novel, druglike ER ligand. Aided by crystal structures of novel ligands bound to an ER construct, medicinal chemistry iterations led to (E)-3-(3,5-difluoro-4-((1R,3R)-2-(2-fluoro-2-methylpropyl)-3-methyl-2,3,4,9-tetrahydro-1H-pyrido[3,4-b]indol-1-yl)phenyl)acrylic acid (30b, AZD9496), a clinical candidate with high oral bioavailability across preclinical species that is currently being evaluated in phase I clinical trials for the treatment of advanced estrogen receptor (ER) positive breast cancer.
Organizational Affiliation:
Oncology iMed, AstraZeneca, Mereside, Alderley Park, Macclesfield SK10 4TG, U.K.