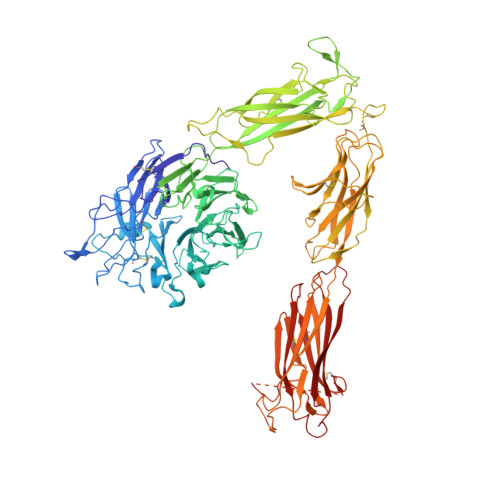
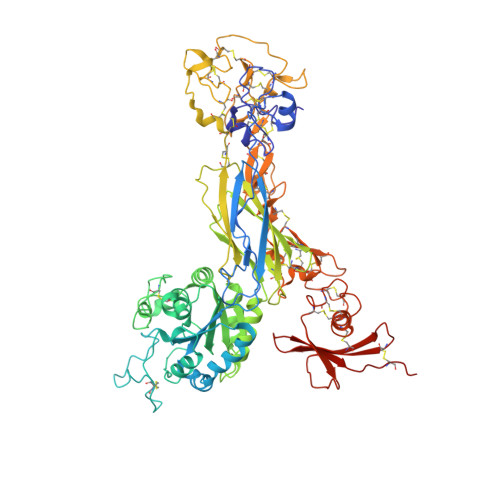
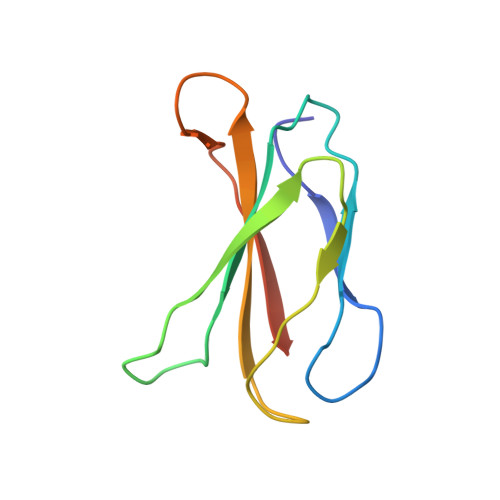
Structural basis for pure antagonism of integrin alpha V beta 3 by a high-affinity form of fibronectin.
Van Agthoven, J.F., Xiong, J.P., Alonso, J.L., Rui, X., Adair, B.D., Goodman, S.L., Arnaout, M.A.(2014) Nat Struct Mol Biol 21: 383-388
- PubMed: 24658351
- DOI: https://doi.org/10.1038/nsmb.2797
- Primary Citation of Related Structures:
4MMX, 4MMY, 4MMZ - PubMed Abstract:
Integrins are important therapeutic targets. However, current RGD-based anti-integrin drugs are also partial agonists, inducing conformational changes that trigger potentially fatal immune reactions and paradoxical cell adhesion. Here we describe the first crystal structure of αVβ3 bound to a physiologic ligand, the tenth type III RGD domain of wild-type fibronectin (wtFN10), or to a high-affinity mutant (hFN10) shown here to act as a pure antagonist. Comparison of these structures revealed a central π-π interaction between Trp1496 in the RGD-containing loop of hFN10 and Tyr122 of the β3 subunit that blocked conformational changes triggered by wtFN10 and trapped hFN10-bound αVβ3 in an inactive conformation. Removing the Trp1496 or Tyr122 side chains or reorienting Trp1496 away from Tyr122 converted hFN10 into a partial agonist. These findings offer new insights into the mechanism of integrin activation and a basis for the design of RGD-based pure antagonists.
Organizational Affiliation:
1] Structural Biology Program, Department of Medicine, Harvard Medical School, Massachusetts General Hospital, Charlestown, Massachusetts, USA. [2].