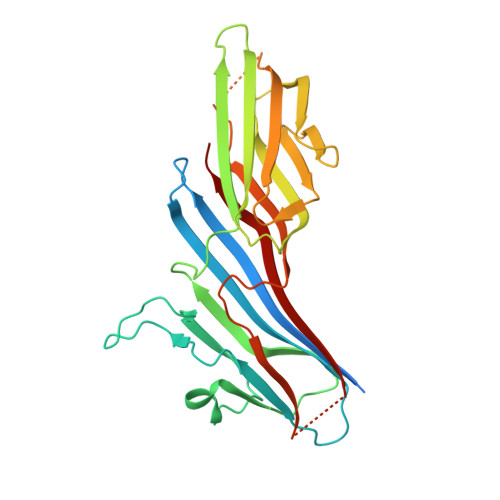
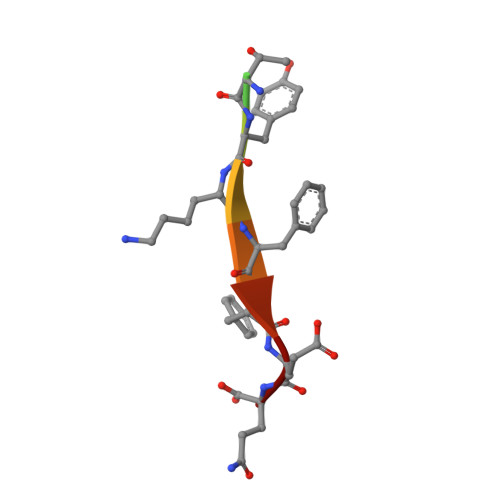
Structural and Functional Characterization of Cargo-Binding Sites on the mu 4-Subunit of Adaptor Protein Complex 4.
Ross, B.H., Lin, Y., Corales, E.A., Burgos, P.V., Mardones, G.A.(2014) PLoS One 9: e88147-e88147
- PubMed: 24498434
- DOI: https://doi.org/10.1371/journal.pone.0088147
- Primary Citation of Related Structures:
4MDR - PubMed Abstract:
Adaptor protein (AP) complexes facilitate protein trafficking by playing key roles in the selection of cargo molecules to be sorted in post-Golgi compartments. Four AP complexes (AP-1 to AP-4) contain a medium-sized subunit (μ1-μ4) that recognizes YXXØ-sequences (Ø is a bulky hydrophobic residue), which are sorting signals in transmembrane proteins. A conserved, canonical region in μ subunits mediates recognition of YXXØ-signals by means of a critical aspartic acid. Recently we found that a non-canonical YXXØ-signal on the cytosolic tail of the Alzheimer's disease amyloid precursor protein (APP) binds to a distinct region of the μ4 subunit of the AP-4 complex. In this study we aimed to determine the functionality of both binding sites of μ4 on the recognition of the non-canonical YXXØ-signal of APP. We found that substitutions in either binding site abrogated the interaction with the APP-tail in yeast-two hybrid experiments. Further characterization by isothermal titration calorimetry showed instead loss of binding to the APP signal with only the substitution R283D at the non-canonical site, in contrast to a decrease in binding affinity with the substitution D190A at the canonical site. We solved the crystal structure of the C-terminal domain of the D190A mutant bound to this non-canonical YXXØ-signal. This structure showed no significant difference compared to that of wild-type μ4. Both differential scanning fluorimetry and limited proteolysis analyses demonstrated that the D190A substitution rendered μ4 less stable, suggesting an explanation for its lower binding affinity to the APP signal. Finally, in contrast to overexpression of the D190A mutant, and acting in a dominant-negative manner, overexpression of μ4 with either a F255A or a R283D substitution at the non-canonical site halted APP transport at the Golgi apparatus. Together, our analyses support that the functional recognition of the non-canonical YXXØ-signal of APP is limited to the non-canonical site of μ4.
Organizational Affiliation:
Instituto de Fisiología, Facultad de Medicina, and Centro de Investigación Sur-Austral en Enfermedades del Sistema Nervioso, Universidad Austral de Chile, Valdivia, Chile.