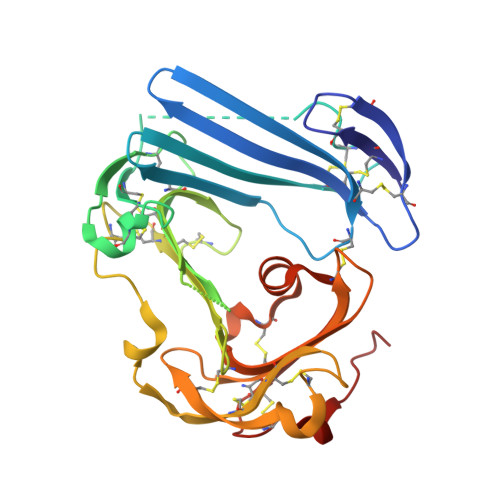
Crystal structure of the urokinase receptor in a ligand-free form.
Xu, X., Gardsvoll, H., Yuan, C., Lin, L., Ploug, M., Huang, M.(2012) J Mol Biol 416: 629-641
- PubMed: 22285761
- DOI: https://doi.org/10.1016/j.jmb.2011.12.058
- Primary Citation of Related Structures:
3U73, 3U74 - PubMed Abstract:
The urokinase receptor urokinase-type plasminogen activator receptor (uPAR) is a surface receptor capable of not only focalizing urokinase-type plasminogen activator (uPA)-mediated fibrinolysis to the pericellular micro-environment but also promoting cell migration and chemotaxis. Consistent with this multifunctional role, uPAR binds several extracellular ligands, including uPA and vitronectin. Structural studies suggest that uPAR possesses structural flexibility. It is, however, not clear whether this flexibility is an inherent property of the uPAR structure per se or whether it is induced upon ligand binding. The crystal structure of human uPAR in its ligand-free state would clarify this issue, but such information remains unfortunately elusive. We now report the crystal structures of a stabilized, human uPAR (H47C/N259C) in its ligand-free form to 2.4 Å and in complex with amino-terminal fragment (ATF) to 3.2 Å. The structure of uPAR(H47C/N259C) in complex with ATF resembles the wild-type uPAR·ATF complex, demonstrating that these mutations do not perturb the uPA binding properties of uPAR. The present structure of uPAR(H47C/N259C) provides the first structural definition of uPAR in its ligand-free form, which represents one of the biologically active conformations of uPAR as defined by extensive biochemical studies. The domain boundary between uPAR DI-DII domains is more flexible than the DII-DIII domain boundary. Two important structural features are highlighted by the present uPAR structure. First, the DI-DIII domain boundary may face the cell membrane. Second, loop 130-140 of uPAR plays a dynamic role during ligand loading/unloading. Together, these studies provide new insights into uPAR structure-function relationships, emphasizing the importance of the inter-domain dynamics of this modular receptor.
Organizational Affiliation:
Beth Israel Deaconess Medical Center, Harvard Medical School, Boston, MA 02215, USA.