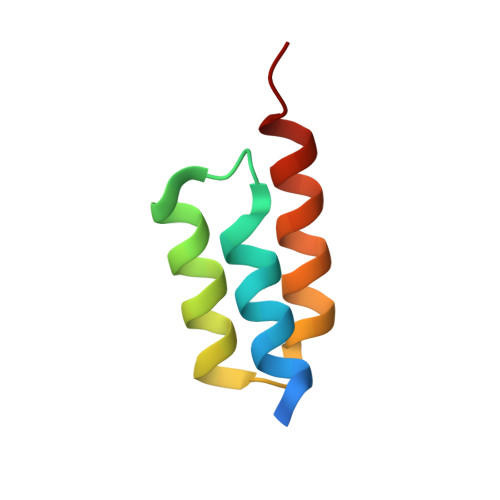
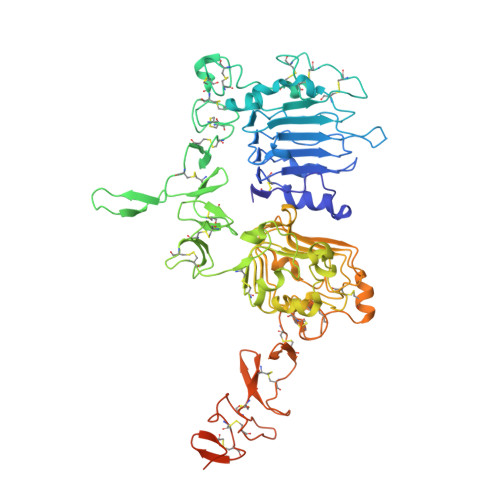
Structural basis for high-affinity HER2 receptor binding by an engineered protein.
Eigenbrot, C., Ultsch, M., Dubnovitsky, A., Abrahmsen, L., Hard, T.(2010) Proc Natl Acad Sci U S A 107: 15039-15044
- PubMed: 20696930
- DOI: https://doi.org/10.1073/pnas.1005025107
- Primary Citation of Related Structures:
2KZI, 2KZJ, 3MZW - PubMed Abstract:
The human epidermal growth factor receptor 2 (HER2) is specifically overexpressed in tumors of several cancers, including an aggressive form of breast cancer. It is therefore a target for both cancer diagnostics and therapy. The 58 amino acid residue Zher2 affibody molecule was previously engineered as a high-affinity binder of HER2. Here we determined the structure of Zher2 in solution and the crystal structure of Zher2 in complex with the HER2 extracellular domain. Zher2 binds to a conformational epitope on HER2 that is distant from those recognized by the therapeutic antibodies trastuzumab and pertuzumab. Its small size and lack of interference may provide Zher2 with advantages for diagnostic use or even for delivery of therapeutic agents to HER2-expressing tumors when trastuzumab or pertuzumab are already employed. Biophysical characterization shows that Zher2 is thermodynamically stable in the folded state yet undergoing conformational interconversion on a submillisecond time scale. The data suggest that it is the HER2-binding conformation that is formed transiently prior to binding. Still, binding is very strong with a dissociation constant K(D) = 22 pM, and perfect conformational homogeneity is therefore not necessarily required in engineered binding proteins. A comparison of the original Z domain scaffold to free and bound Zher2 structures reveals how high-affinity binding has evolved during selection and affinity maturation and suggests how a compromise between binding surface optimization and stability and dynamics of the unbound state has been reached.
Organizational Affiliation:
Department of Structural Biology, Genentech Inc, 1 DNA Way, South San Francisco, CA 94080, USA. eigenbrot.c@gene.com