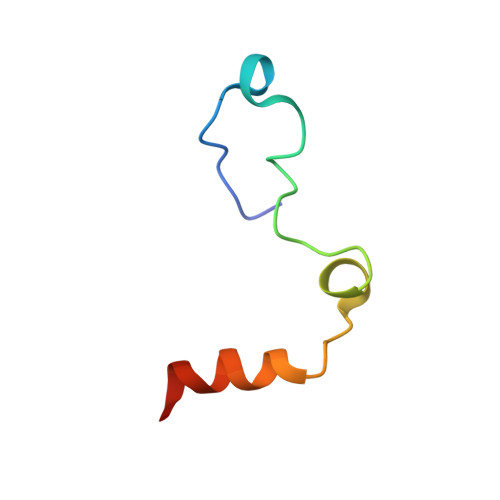
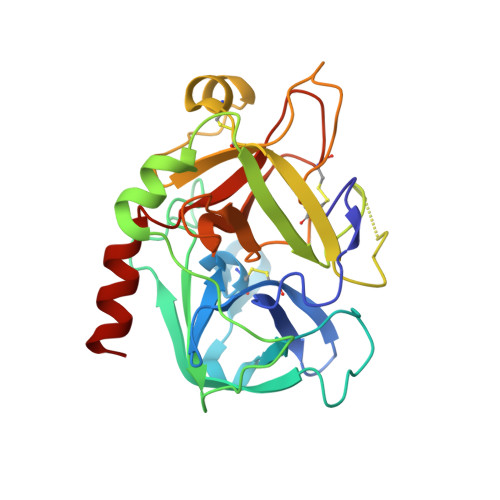
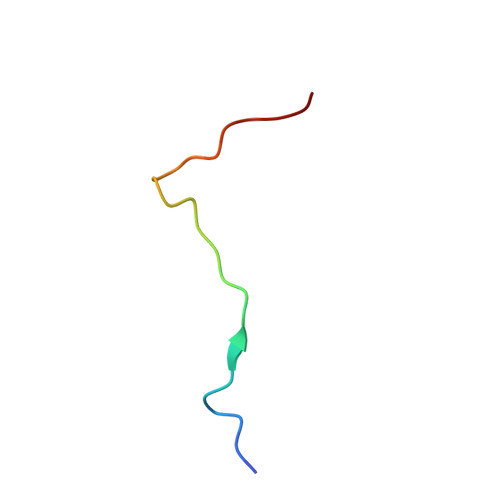
Crystal structure of thrombin bound to the uncleaved extracellular fragment of PAR1.
Gandhi, P.S., Chen, Z., Di Cera, E.(2010) J Biol Chem 285: 15393-15398
- PubMed: 20236938
- DOI: https://doi.org/10.1074/jbc.M110.115337
- Primary Citation of Related Structures:
3LU9 - PubMed Abstract:
Abundant structural information exists on how thrombin recognizes ligands at the active site or at exosites separate from the active site region, but remarkably little is known about how thrombin recognizes substrates that bridge both the active site and exosite I. The case of the protease-activated receptor PAR1 is particularly relevant in view of the plethora of biological effects associated with its activation by thrombin. Here, we present the 1.8 A resolution structure of thrombin S195A in complex with a 30-residue long uncleaved extracellular fragment of PAR1 that documents for the first time a productive binding mode bridging the active site and exosite I. The structure reveals two unexpected features of the thrombin-PAR1 interaction. The acidic P3 residue of PAR1, Asp(39), does not hinder binding to the active site and actually makes favorable interactions with Gly(219) of thrombin. The tethered ligand domain shows a considerable degree of disorder even when bound to thrombin. The results fill a significant gap in our understanding of the molecular mechanisms of recognition by thrombin in ways that are relevant to other physiological substrates.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Saint Louis University School of Medicine, St. Louis, Missouri 63104.