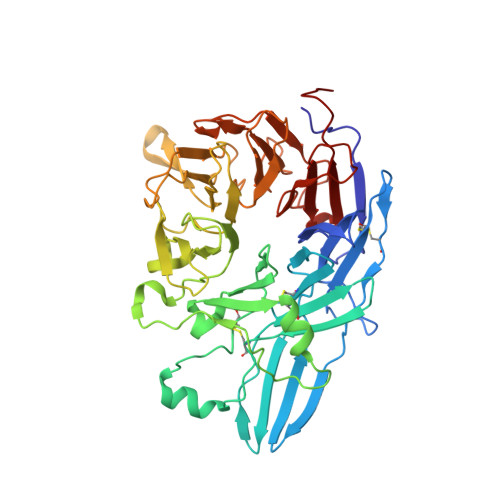
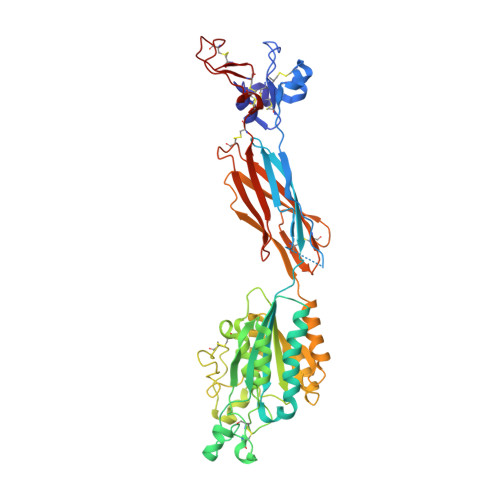
Structure of a complete integrin ectodomain in a physiologic resting state and activation and deactivation by applied forces.
Zhu, J., Luo, B.H., Xiao, T., Zhang, C., Nishida, N., Springer, T.A.(2008) Mol Cell 32: 849-861
- PubMed: 19111664
- DOI: https://doi.org/10.1016/j.molcel.2008.11.018
- Primary Citation of Related Structures:
3FCS, 3FCU - PubMed Abstract:
The complete ectodomain of integrin alpha(IIb)beta(3) reveals a bent, closed, low-affinity conformation, the beta knee, and a mechanism for linking cytoskeleton attachment to high affinity for ligand. Ca and Mg ions in the recognition site, including the synergistic metal ion binding site (SyMBS), are loaded prior to ligand binding. Electrophilicity of the ligand-binding Mg ion is increased in the open conformation. The beta(3) knee passes between the beta(3)-PSI and alpha(IIb)-knob to bury the lower beta leg in a cleft, from which it is released for extension. Different integrin molecules in crystals and EM reveal breathing that appears on pathway to extension. Tensile force applied to the extended ligand-receptor complex stabilizes the closed, low-affinity conformation. By contrast, an additional lateral force applied to the beta subunit to mimic attachment to moving actin filaments stabilizes the open, high-affinity conformation. This mechanism propagates allostery over long distances and couples cytoskeleton attachment of integrins to their high-affinity state.
Organizational Affiliation:
The Immune Disease Institute and Department of Pathology, Harvard Medical School, 200 Longwood Avenue, Boston, MA 02115, USA.