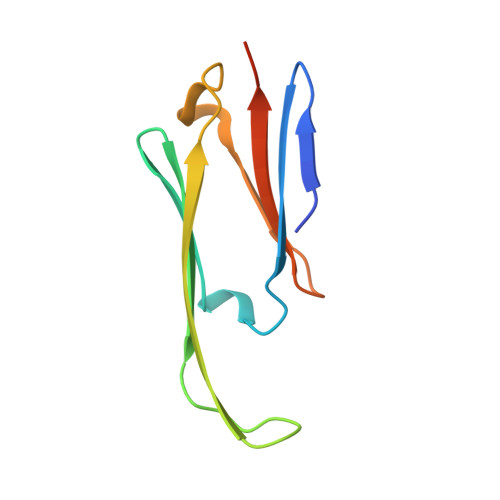
Crystal Structures of Alpha-Crystallin Domain Dimers of Alphab-Crystallin and Hsp20.
Bagneris, C., Bateman, O.A., Naylor, C.E., Cronin, N., Boelens, W.C., Keep, N.H., Slingsby, C.(2009) J Mol Biol 392: 1242
- PubMed: 19646995
- DOI: https://doi.org/10.1016/j.jmb.2009.07.069
- Primary Citation of Related Structures:
2WJ5, 2WJ7 - PubMed Abstract:
Small heat shock proteins (sHsps) are a family of large and dynamic oligomers highly expressed in long-lived cells of muscle, lens and brain. Several family members are upregulated during stress, and some are strongly cytoprotective. Their polydispersity has hindered high-resolution structure analyses, particularly for vertebrate sHsps. Here, crystal structures of excised alpha-crystallin domain from rat Hsp20 and that from human alphaB-crystallin show that they form homodimers with a shared groove at the interface by extending a beta sheet. However, the two dimers differ in the register of their interfaces. The dimers have empty pockets that in large assemblies will likely be filled by hydrophobic sequence motifs from partner chains. In the Hsp20 dimer, the shared groove is partially filled by peptide in polyproline II conformation. Structural homology with other sHsp crystal structures indicates that in full-length chains the groove is likely filled by an N-terminal extension. Inside the groove is a symmetry-related functionally important arginine that is mutated, or its equivalent, in family members in a range of neuromuscular diseases and cataract. Analyses of residues within the groove of the alphaB-crystallin interface show that it has a high density of positive charges. The disease mutant R120G alpha-crystallin domain dimer was found to be more stable at acidic pH, suggesting that the mutation affects the normal dynamics of sHsp assembly. The structures provide a starting point for modelling higher assembly by defining the spatial locations of grooves and pockets in a basic dimeric assembly unit. The structures provide a high-resolution view of a candidate functional state of an sHsp that could bind non-native client proteins or specific components from cytoprotective pathways. The empty pockets and groove provide a starting model for designing drugs to inhibit those sHsps that have a negative effect on cancer treatment.
Organizational Affiliation:
Department of Crystallography, Birkbeck College, Institute of Structural and Molecular Biology, Malet Street, London WC1E 7HX, UK.