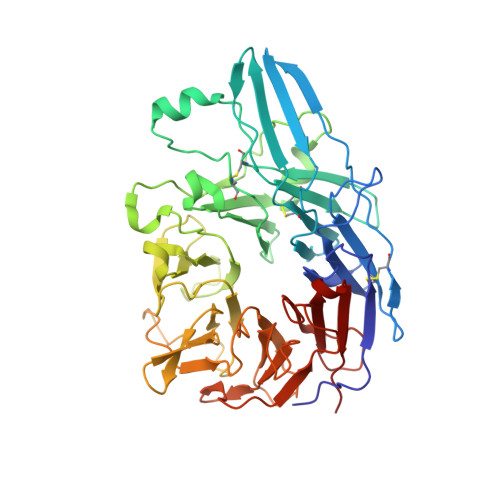
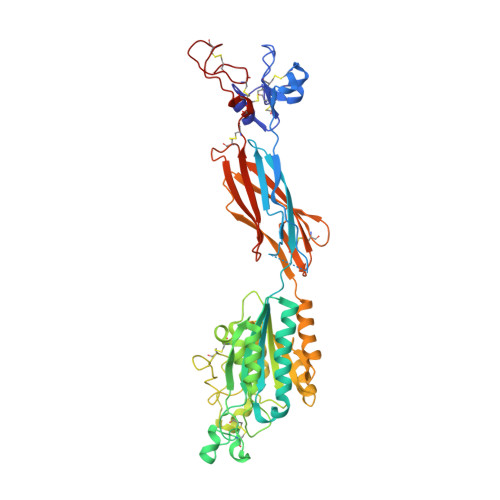
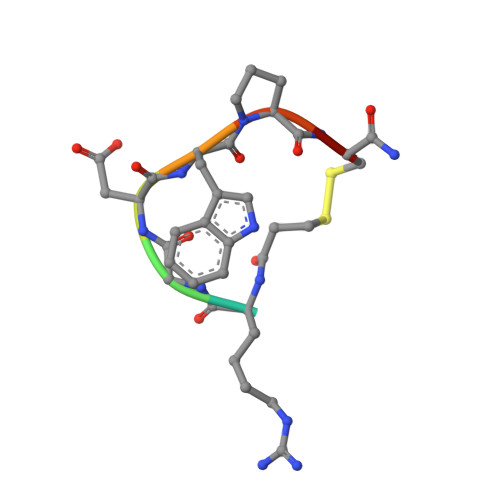
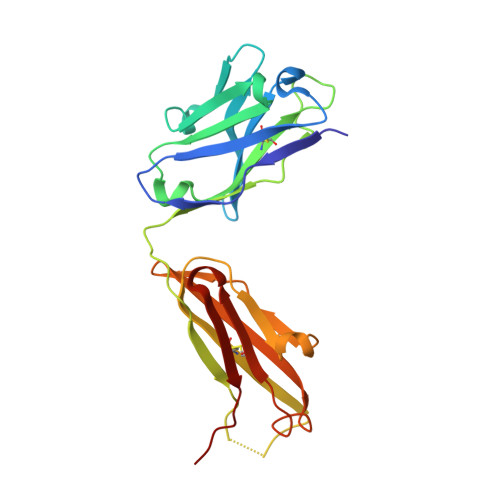
Structural Basis for Distinctive Recognition of Fibrinogen Gammac Peptide by the Platelet Integrin Alphaiibbeta3.
Springer, T.A., Zhu, J., Xiao, T.(2008) J Cell Biol 182: 791
- PubMed: 18710925
- DOI: https://doi.org/10.1083/jcb.200801146
- Primary Citation of Related Structures:
2VC2, 2VDK, 2VDL, 2VDM, 2VDN, 2VDO, 2VDP, 2VDQ, 2VDR - PubMed Abstract:
Hemostasis and thrombosis (blood clotting) involve fibrinogen binding to integrin alpha(IIb)beta(3) on platelets, resulting in platelet aggregation. alpha(v)beta(3) binds fibrinogen via an Arg-Asp-Gly (RGD) motif in fibrinogen's alpha subunit. alpha(IIb)beta(3) also binds to fibrinogen; however, it does so via an unstructured RGD-lacking C-terminal region of the gamma subunit (gammaC peptide). These distinct modes of fibrinogen binding enable alpha(IIb)beta(3) and alpha(v)beta(3) to function cooperatively in hemostasis. In this study, crystal structures reveal the integrin alpha(IIb)beta(3)-gammaC peptide interface, and, for comparison, integrin alpha(IIb)beta(3) bound to a lamprey gammaC primordial RGD motif. Compared with RGD, the GAKQAGDV motif in gammaC adopts a different backbone configuration and binds over a more extended region. The integrin metal ion-dependent adhesion site (MIDAS) Mg(2+) ion binds the gammaC Asp side chain. The adjacent to MIDAS (ADMIDAS) Ca(2+) ion binds the gammaC C terminus, revealing a contribution for ADMIDAS in ligand binding. Structural data from this natively disordered gammaC peptide enhances our understanding of the involvement of gammaC peptide and integrin alpha(IIb)beta(3) in hemostasis and thrombosis.
Organizational Affiliation:
Department of Pathology, Immune Disease Institute, Harvard Medical School, Boston, MA 02115, USA.