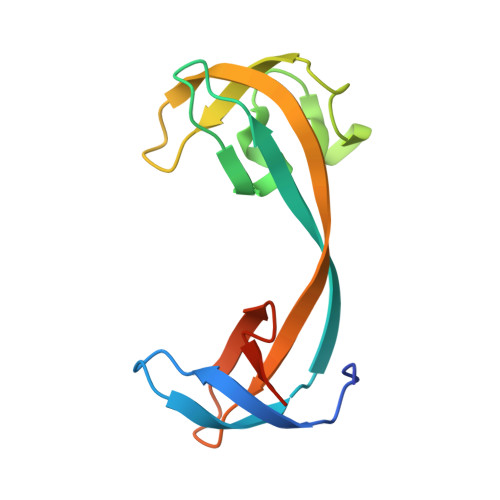
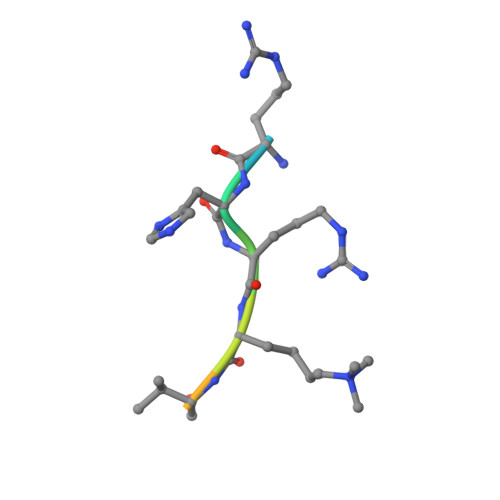
Distinct binding modes specify the recognition of methylated histones H3K4 and H4K20 by JMJD2A-tudor.
Lee, J., Thompson, J.R., Botuyan, M.V., Mer, G.(2008) Nat Struct Mol Biol 15: 109-111
- PubMed: 18084306
- DOI: https://doi.org/10.1038/nsmb1326
- Primary Citation of Related Structures:
2QQR, 2QQS - PubMed Abstract:
The lysine demethylase JMJD2A has the unique property of binding trimethylated peptides from two different histone sequences (H3K4me3 and H4K20me3) through its tudor domains. Here we show using X-ray crystallography and calorimetry that H3K4me3 and H4K20me3, which are recognized with similar affinities by JMJD2A, adopt radically different binding modes, to the extent that we were able to design single point mutations in JMJD2A that inhibited the recognition of H3K4me3 but not H4K20me3 and vice versa.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Mayo Clinic College of Medicine, Rochester, Minnesota 55905, USA.