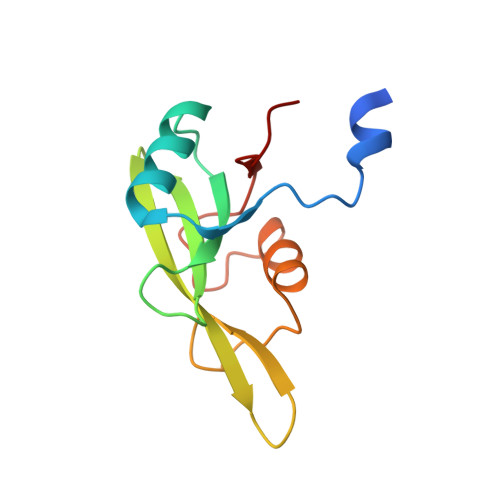
Grb7 SH2 domain structure and interactions with a cyclic peptide inhibitor of cancer cell migration and proliferation.
Porter, C.J., Matthews, J.M., Mackay, J.P., Pursglove, S.E., Schmidberger, J.W., Leedman, P.J., Pero, S.C., Krag, D.N., Wilce, M.C., Wilce, J.A.(2007) BMC Struct Biol 7: 58-58
- PubMed: 17894853
- DOI: https://doi.org/10.1186/1472-6807-7-58
- Primary Citation of Related Structures:
2QMS - PubMed Abstract:
Human growth factor receptor bound protein 7 (Grb7) is an adapter protein that mediates the coupling of tyrosine kinases with their downstream signaling pathways. Grb7 is frequently overexpressed in invasive and metastatic human cancers and is implicated in cancer progression via its interaction with the ErbB2 receptor and focal adhesion kinase (FAK) that play critical roles in cell proliferation and migration. It is thus a prime target for the development of novel anti-cancer therapies. Recently, an inhibitory peptide (G7-18NATE) has been developed which binds specifically to the Grb7 SH2 domain and is able to attenuate cancer cell proliferation and migration in various cancer cell lines. As a first step towards understanding how Grb7 may be inhibited by G7-18NATE, we solved the crystal structure of the Grb7 SH2 domain to 2.1 A resolution. We describe the details of the peptide binding site underlying target specificity, as well as the dimer interface of Grb 7 SH2. Dimer formation of Grb7 was determined to be in the muM range using analytical ultracentrifugation for both full-length Grb7 and the SH2 domain alone, suggesting the SH2 domain forms the basis of a physiological dimer. ITC measurements of the interaction of the G7-18NATE peptide with the Grb7 SH2 domain revealed that it binds with a binding affinity of Kd = approximately 35.7 microM and NMR spectroscopy titration experiments revealed that peptide binding causes perturbations to both the ligand binding surface of the Grb7 SH2 domain as well as to the dimer interface, suggesting that dimerisation of Grb7 is impacted on by peptide binding. Together the data allow us to propose a model of the Grb7 SH2 domain/G7-18NATE interaction and to rationalize the basis for the observed binding specificity and affinity. We propose that the current study will assist with the development of second generation Grb7 SH2 domain inhibitors, potentially leading to novel inhibitors of cancer cell migration and invasion.
Organizational Affiliation:
School of Biomedical and Chemical Sciences, University of Western Australia, WA 6009, Australia. Corrine.Porter@med.monash.edu.au