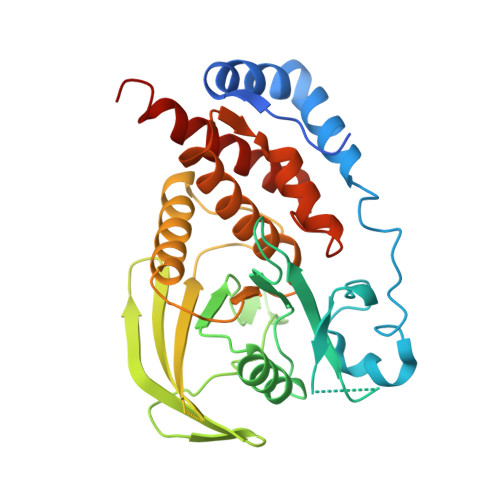
Engineering the catalytic domain of human protein tyrosine phosphatase beta for structure-based drug discovery.
Evdokimov, A.G., Pokross, M., Walter, R., Mekel, M., Cox, B., Li, C., Bechard, R., Genbauffe, F., Andrews, R., Diven, C., Howard, B., Rastogi, V., Gray, J., Maier, M., Peters, K.G.(2006) Acta Crystallogr D Biol Crystallogr 62: 1435-1445
- PubMed: 17139078
- DOI: https://doi.org/10.1107/S0907444906037784
- Primary Citation of Related Structures:
2HC1, 2HC2, 2I3R, 2I3U, 2I4E, 2I4G, 2I4H, 2I5X - PubMed Abstract:
Protein tyrosine phosphatases (PTPs) play roles in many biological processes and are considered to be important targets for drug discovery. As inhibitor development has proven challenging, crystal structure-based design will be very helpful to advance inhibitor potency and selectivity. Successful application of protein crystallography to drug discovery heavily relies on high-quality crystal structures of the protein of interest complexed with pharmaceutically interesting ligands. It is very important to be able to produce protein-ligand crystals rapidly and reproducibly for as many ligands as necessary. This study details our efforts to engineer the catalytic domain of human protein tyrosine phosphatase beta (HPTPbeta-CD) with properties suitable for rapid-turnaround crystallography. Structures of apo HPTPbeta-CD and its complexes with several novel small-molecule inhibitors are presented here for the first time.
Organizational Affiliation:
Structural Biology, Procter and Gamble Pharmaceuticals, 8700 Mason-Montgomery Road, Mason, OH 45140, USA. artem@xtals.org