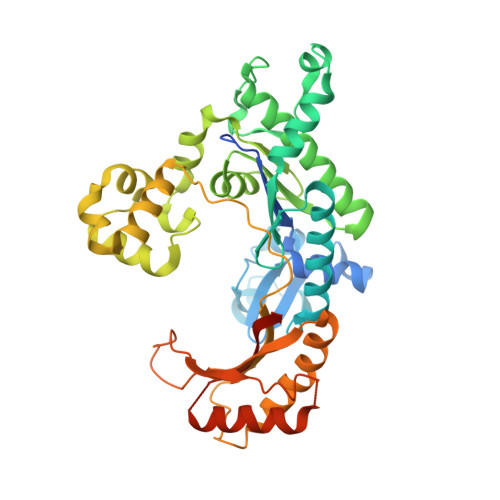
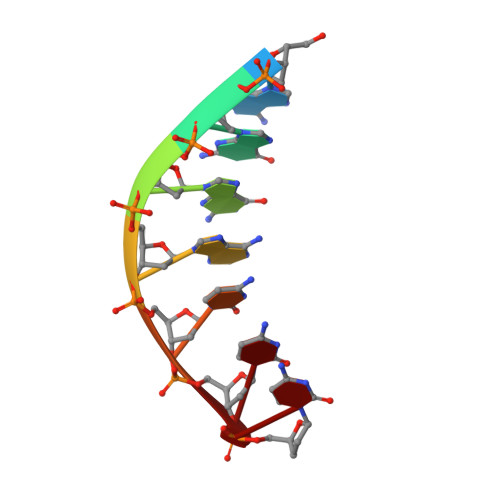
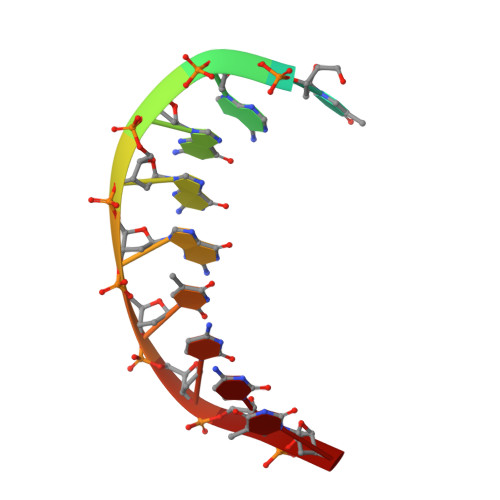
An incoming nucleotide imposes an anti to syn conformational change on the templating purine in the human DNA polymerase-iota active site.
Nair, D.T., Johnson, R.E., Prakash, L., Prakash, S., Aggarwal, A.K.(2006) Structure 14: 749-755
- PubMed: 16615915
- DOI: https://doi.org/10.1016/j.str.2006.01.010
- Primary Citation of Related Structures:
2FLL, 2FLN, 2FLP - PubMed Abstract:
Substrate-induced conformational change of the protein is the linchpin of enzymatic reactions. Replicative DNA polymerases, for example, convert from an open to a closed conformation in response to dNTP binding. Human DNA polymerase-iota (hPoliota), a member of the Y family of DNA polymerases, differs strikingly from other polymerases in its much higher proficiency and fidelity for nucleotide incorporation opposite template purines than opposite template pyrimidines. We present here a crystallographic analysis of hPoliota binary complexes, which together with the ternary complexes show that, contrary to replicative DNA polymerases, the DNA, and not the polymerase, undergoes the primary substrate-induced conformational change. The incoming dNTP "pushes" templates A and G from the anti to the syn conformation dictated by a rigid hPoliota active site. Together, the structures posit a mechanism for template selection wherein dNTP binding induces a conformational switch in template purines for productive Hoogsteen base pairing.
Organizational Affiliation:
Structural Biology Program, Department of Physiology and Biophysics, Mount Sinai School of Medicine, Box 1677, 1425 Madison Avenue, New York, New York 10029, USA.