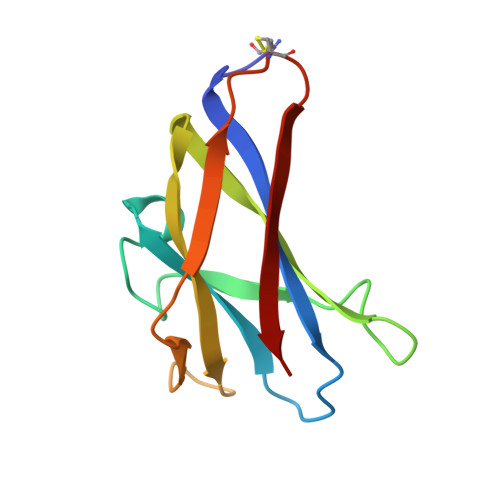
Solution structure of the granular starch binding domain of glucoamylase from Aspergillus niger by nuclear magnetic resonance spectroscopy.
Sorimachi, K., Jacks, A.J., Le Gal-Coeffet, M.F., Williamson, G., Archer, D.B., Williamson, M.P.(1996) J Mol Biol 259: 970-987
- PubMed: 8683599
- DOI: https://doi.org/10.1006/jmbi.1996.0374
- Primary Citation of Related Structures:
1KUL, 1KUM - PubMed Abstract:
The solution structure of the granular starch binding domain (SBD) of glucoamylase 1 from Aspergillus niger has been determined by heteronuclear multidimensional nuclear magnetic resonance spectroscopy and simulated annealing. A total of 1092 nuclear Overhauser enhancement-derived 1H-1H distance constraints, 137 dihedral constraints and 86 hydrogen bond constraints were incorporated into an X-PLOR simulated annealing and refinement protocol. The family of calculated structures shows a well defined beta-sheet structure consisting of one parallel and six antiparallel pairs of beta-strands which forms an open-sided beta-barrel. The root-mean-square deviation (rmsd) of 53 individual structures to the calculated average structure for the backbone atoms of residues excluding the N terminus and two mobile loops is 0.57(+/-0.10) A while the rmsd for backbone atoms in beta-strands is 0.45(+/-0.08) A. Structural features of the SBD in solution are compared to the X-ray crystal structure of a homologous domain of cyclodextrin glycosyltransferase (CGTase) in the free and bound forms. Titration studies with two ligands, maltoheptaose and beta-cyclodextrin, show the existence of two binding sites. Examination of the tertiary structures shows these two sites to be at one end of the molecule on opposite faces. The majority of residues showing the largest 1H and 15N chemical shift changes are located in loop regions. Many residues implicated in binding, based on these changes, are similar in location to previously identified binding site residues in the crystal structures of CGTase. Overall, the shift changes are small indicating that the SBD does not undergo large conformational changes upon ligand binding.
Organizational Affiliation:
Krebs Institute for Biomolecular Research, Department of Molecular Biology and Biotechnology, University of Sheffield, UK.