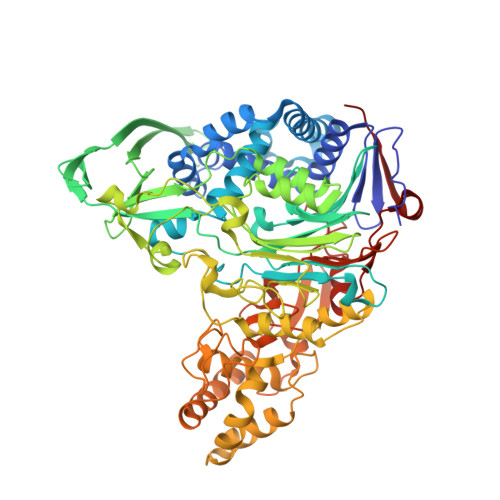
Precursor structure of cephalosporin acylase. Insights into autoproteolytic activation in a new N-terminal hydrolase family
Kim, Y., Kim, S., Earnest, T.N., Hol, W.G.(2002) J Biol Chem 277: 2823-2829
- PubMed: 11706000
- DOI: https://doi.org/10.1074/jbc.M108888200
- Primary Citation of Related Structures:
1KEH - PubMed Abstract:
Autocatalytic proteolytic cleavage is a frequently observed post-translational modification in proteins. Cephalosporin acylase (CA) is a recently identified member of the N-terminal hydrolase family that is activated from an inactive precursor by autoproteolytic processing, generating a new N-terminal residue, which is either a Ser or a Thr. The N-terminal Ser or Thr becomes a nucleophilic catalytic center for intramolecular and intermolecular amide cleavages. The gene structure of the open reading frame of CAs generally consists of a signal peptide followed by the alpha-subunit, a spacer sequence, and the beta-subunit, which are all translated into a single polypeptide chain, the CA precursor. The precursor is post-translationally modified into an active heterodimeric enzyme with alpha- and beta-subunits, first by intramolecular cleavage and second by intermolecular cleavage. We solved the first CA precursor structure (code 1KEH) from a class I CA from Pseudomonas diminuta at a 2.5-A resolution that provides insight into the mechanism of intramolecular cleavage. A conserved water molecule, stabilized by four hydrogen bonds in unusual pseudotetrahedral geometry, plays a key role to assist the OG atom of Ser(1beta) to generate a strong nucleophile. In addition, the site of the secondary intermolecular cleavage of CA is proposed to be the carbonyl carbon of Gly(158alpha) (Kim, S., and Kim, Y., (2001) J. Biol. Chem., 276, 48376-48381), which is different from the situation in two other class I CAs.
Organizational Affiliation:
School of Chemical Engineering, Yeungnam University, Dae-Dong, Kyungsan 712-749, Korea. ykim1@yu.ac.kr