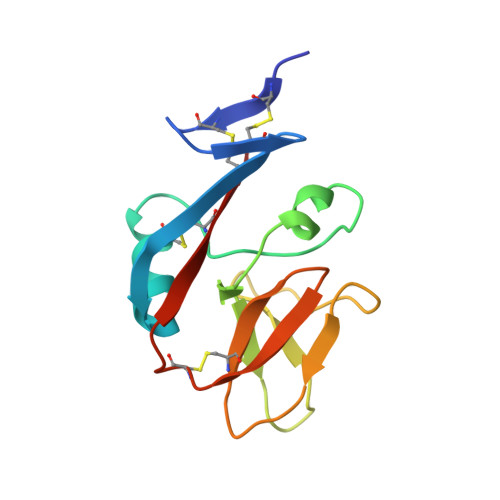
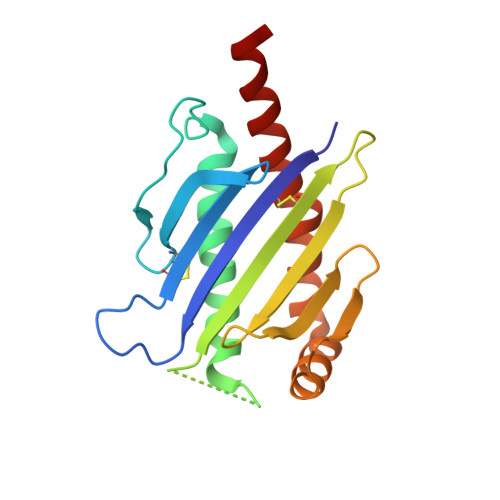
Conformational plasticity revealed by the cocrystal structure of NKG2D and its class I MHC-like ligand ULBP3.
Radaev, S., Rostro, B., Brooks, A.G., Colonna, M., Sun, P.D.(2001) Immunity 15: 1039-1049
- PubMed: 11754823
- DOI: https://doi.org/10.1016/s1074-7613(01)00241-2
- Primary Citation of Related Structures:
1KCG - PubMed Abstract:
NKG2D is known to trigger the natural killer (NK) cell lysis of various tumor and virally infected cells. In the NKG2D/ULBP3 complex, the structure of ULBP3 resembles the alpha1 and alpha2 domains of classical MHC molecules without a bound peptide. The lack of alpha3 and beta2m domains is compensated by replacing two hydrophobic patches at the underside of the class I MHC-like beta sheet floor with a group of hydrophilic and charged residues in ULBP3. NKG2D binds diagonally across the ULBP3 alpha helices, creating a complementary interface, an asymmetrical subunit orientation, and local conformational adjustments in the receptor. The interface is stabilized primarily by hydrogen bonds and hydrophobic interactions. Unlike the KIR receptors that recognize a conserved HLA region by a lock-and-key mechanism, NKG2D recognizes diverse ligands by an induced-fit mechanism.
Organizational Affiliation:
Structural Biology Section, Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases, National Institutes of Health, 12441 Parklawn Drive, Rockville, MD 20852, USA.