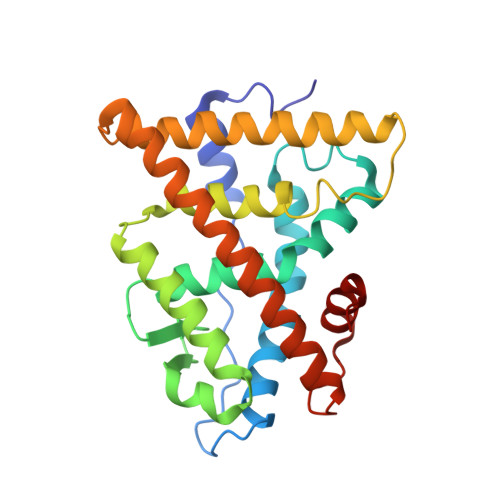
Overexpression, Purification, and Crystal Structure of Native ER alpha LBD
Eiler, S., Gangloff, M., Duclaud, S., Moras, D., Ruff, M.(2001) Protein Expr Purif 22: 165-173
- PubMed: 11437591
- DOI: https://doi.org/10.1006/prep.2001.1409
- Primary Citation of Related Structures:
1G50 - PubMed Abstract:
Several crystal structures of human estrogen receptor alpha ligand-binding domain (hERalpha LBD) complexed with agonist or antagonist molecules have previously been solved. The proteins had been modified in cysteine residues (carboxymethylation) or renatured in urea to circumvent aggregation and denaturation problems. In this work, high-level protein expression and purification together with crystallization screening procedure yielded high amounts of soluble protein without renaturation or modifications steps. The native protein crystallizes in the space group P3(2) 21 with three molecules in the asymmetric unit. The overall structure is very similar to that previously reported for the hERalpha LBD with cysteine carboxymethylated residues thus validating the modification approach. The present strategy can be adapted to other cases where the solubility and the proper folding is a difficulty.
Organizational Affiliation:
Laboratoire de Biologie et Génomique Structurales 1, IGBMC, rue Laurent Fries, Illkirch, 67404, France.