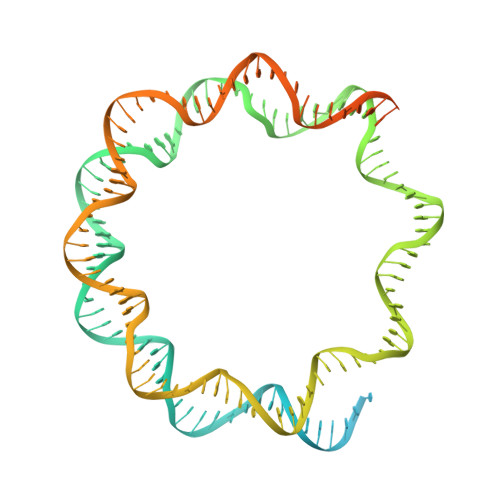
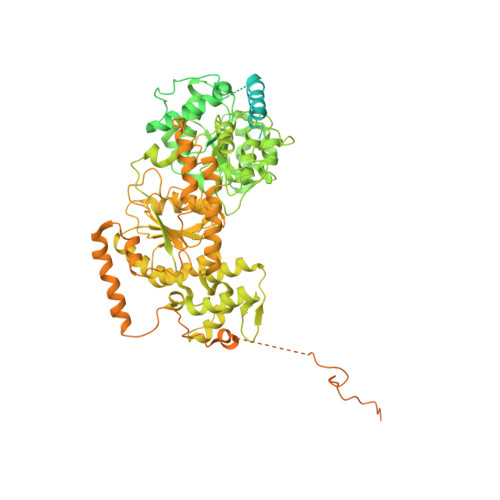
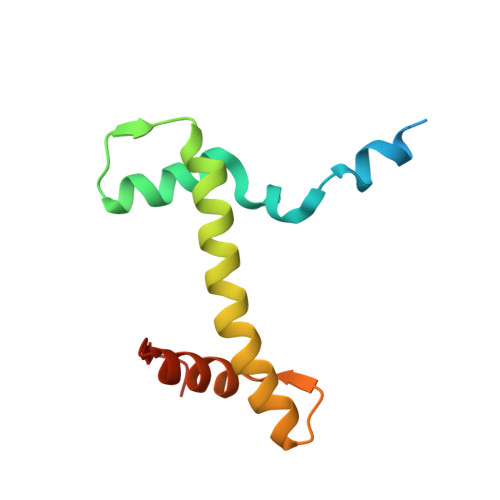
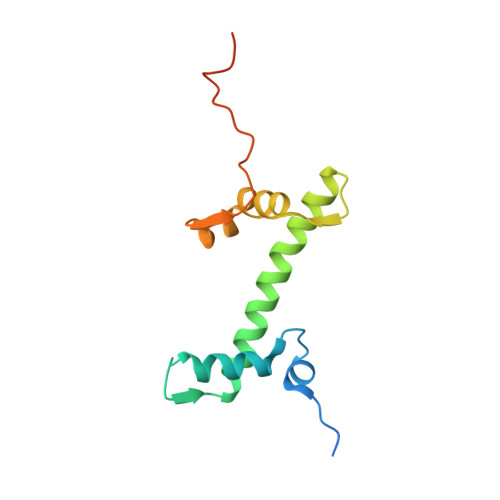
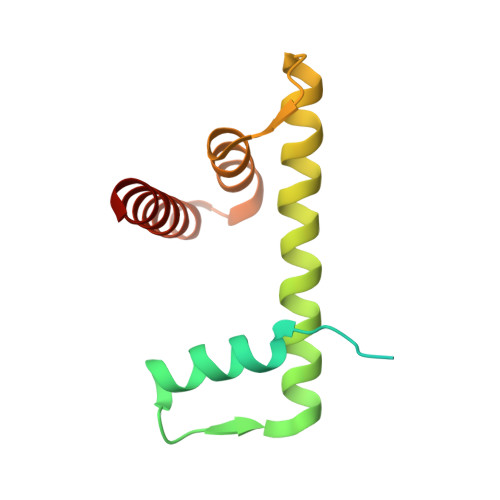
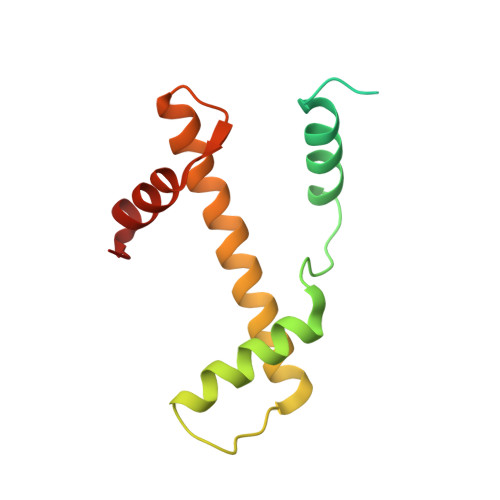
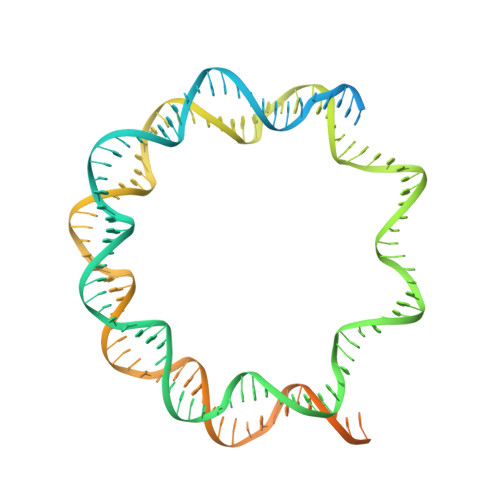
Structure of human chromatin-remodelling PBAF complex bound to a nucleosome.
Yuan, J.J., Chen, K.J., Zhang, W.B., Chen, Z.C.(2022) Nature 605: 166-171
- PubMed: 35477757
- DOI: https://doi.org/10.1038/s41586-022-04658-5
- Primary Citation of Related Structures:
7VDT, 7VDV - PubMed Abstract:
DNA wraps around the histone octamer to form nucleosomes 1 , the repeating unit of chromatin, which create barriers for accessing genetic information. Snf2-like chromatin remodellers couple the energy of ATP binding and hydrolysis to reposition and recompose the nucleosome, and have vital roles in various chromatin-based transactions 2,3 . Here we report the cryo-electron microscopy structure of the 12-subunit human chromatin-remodelling polybromo-associated BRG1-associated factor (PBAF) complex bound to the nucleosome. The motor subunit SMARCA4 engages the nucleosome in the active conformation, which reveals clustering of multiple disease-associated mutations at the interfaces that are essential for chromatin-remodelling activity. SMARCA4 recognizes the H2A-H2B acidic pocket of the nucleosome through three arginine anchors of the Snf2 ATP coupling (SnAc) domain. PBAF shows notable functional modularity, and most of the auxiliary subunits are interwoven into three lobe-like submodules for nucleosome recognition. The PBAF-specific auxiliary subunit ARID2 acts as the structural core for assembly of the DNA-binding lobe, whereas PBRM1, PHF10 and BRD7 are collectively incorporated into the lobe for histone tail binding. Together, our findings provide mechanistic insights into nucleosome recognition by PBAF and a structural basis for understanding SMARCA4-related human diseases.
Organizational Affiliation:
MOE Key Laboratory of Protein Science, Tsinghua University, Beijing, China.