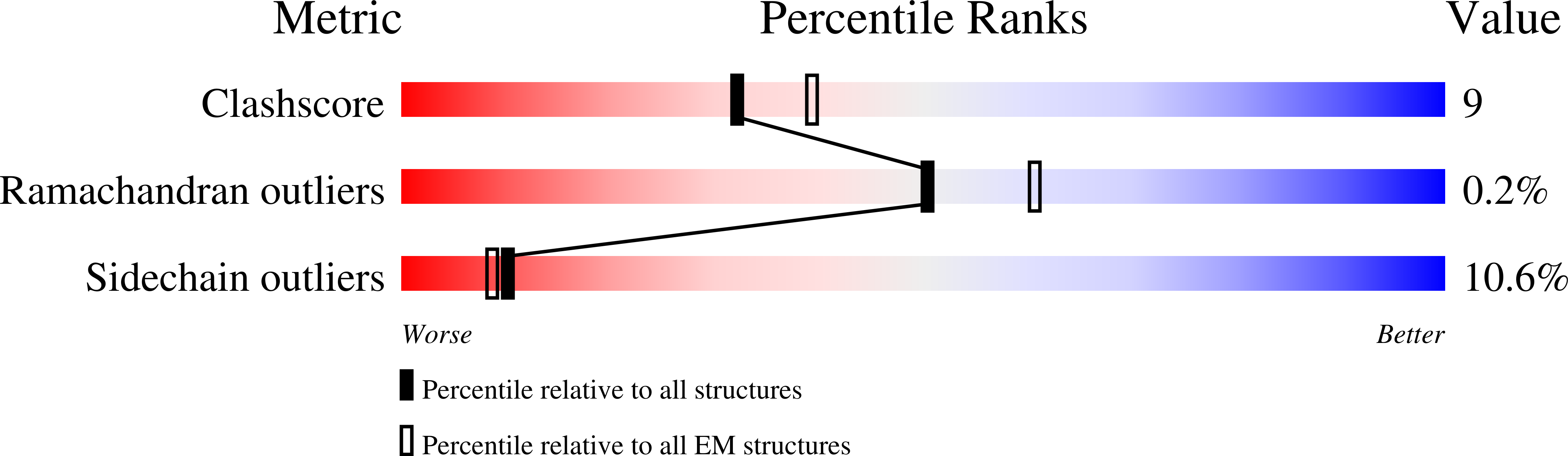Structural basis for transcription activation by Crl through tethering of sigmaSand RNA polymerase.
Cartagena, A.J., Banta, A.B., Sathyan, N., Ross, W., Gourse, R.L., Campbell, E.A., Darst, S.A.(2019) Proc Natl Acad Sci U S A 116: 18923-18927
- PubMed: 31484766
- DOI: https://doi.org/10.1073/pnas.1910827116
- Primary Citation of Related Structures:
6OMF - PubMed Abstract:
In bacteria, a primary σ-factor associates with the core RNA polymerase (RNAP) to control most transcription initiation, while alternative σ-factors are used to coordinate expression of additional regulons in response to environmental conditions. Many alternative σ-factors are negatively regulated by anti-σ-factors. In Escherichia coli , Salmonella enterica , and many other γ-proteobacteria, the transcription factor Crl positively regulates the alternative σ S -regulon by promoting the association of σ S with RNAP without interacting with promoter DNA. The molecular mechanism for Crl activity is unknown. Here, we determined a single-particle cryo-electron microscopy structure of Crl-σ S -RNAP in an open promoter complex with a σ S -regulon promoter. In addition to previously predicted interactions between Crl and domain 2 of σ S (σ S 2 ), the structure, along with p -benzoylphenylalanine cross-linking, reveals that Crl interacts with a structural element of the RNAP β'-subunit that we call the β'-clamp-toe (β'CT). Deletion of the β'CT decreases activation by Crl without affecting basal transcription, highlighting the functional importance of the Crl-β'CT interaction. We conclude that Crl activates σ S -dependent transcription in part through stabilizing σ S -RNAP by tethering σ S 2 and the β'CT. We propose that Crl, and other transcription activators that may use similar mechanisms, be designated σ-activators.
Organizational Affiliation:
Tri-Institutional Training Program in Chemical Biology, The Rockefeller University, New York, NY 10065.