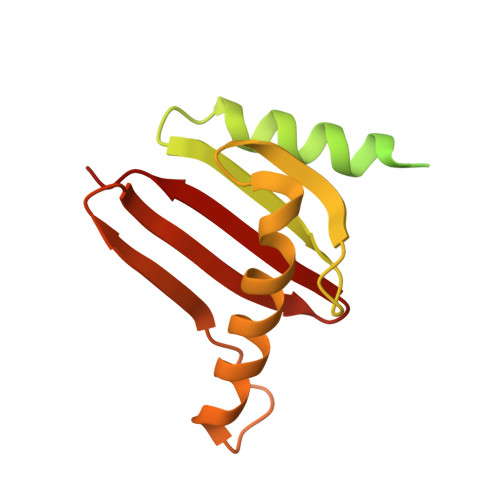
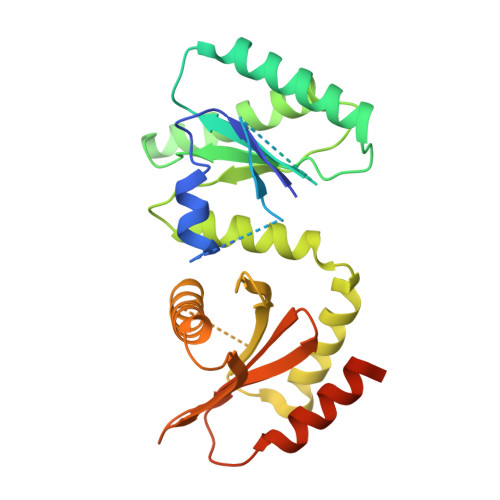
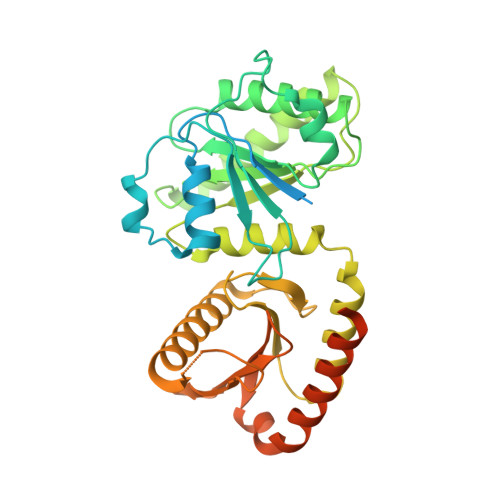
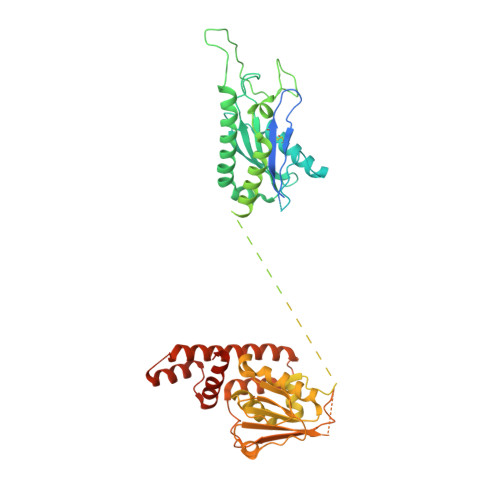
Structural mechanism of a Rag GTPase activation checkpoint by the lysosomal folliculin complex.
Lawrence, R.E., Fromm, S.A., Fu, Y., Yokom, A.L., Kim, D.J., Thelen, A.M., Young, L.N., Lim, C.Y., Samelson, A.J., Hurley, J.H., Zoncu, R.(2019) Science 366: 971-977
- PubMed: 31672913
- DOI: https://doi.org/10.1126/science.aax0364
- Primary Citation of Related Structures:
6NZD - PubMed Abstract:
The tumor suppressor folliculin (FLCN) enables nutrient-dependent activation of the mechanistic target of rapamycin complex 1 (mTORC1) protein kinase via its guanosine triphosphatase (GTPase) activating protein (GAP) activity toward the GTPase RagC. Concomitant with mTORC1 inactivation by starvation, FLCN relocalizes from the cytosol to lysosomes. To determine the lysosomal function of FLCN, we reconstituted the human lysosomal FLCN complex (LFC) containing FLCN, its partner FLCN-interacting protein 2 (FNIP2), and the RagA GDP :RagC GTP GTPases as they exist in the starved state with their lysosomal anchor Ragulator complex and determined its cryo-electron microscopy structure to 3.6 angstroms. The RagC-GAP activity of FLCN was inhibited within the LFC, owing to displacement of a catalytically required arginine in FLCN from the RagC nucleotide. Disassembly of the LFC and release of the RagC-GAP activity of FLCN enabled mTORC1-dependent regulation of the master regulator of lysosomal biogenesis, transcription factor E3, implicating the LFC as a checkpoint in mTORC1 signaling.
Organizational Affiliation:
Department of Molecular and Cell Biology, University of California at Berkeley, Berkeley, CA 94720, USA.