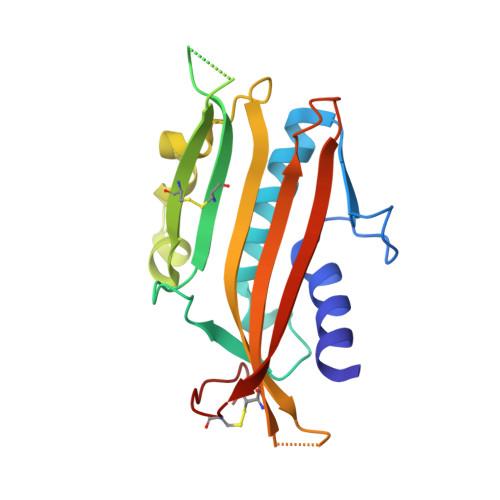
Structure of the Human TRPML2 Ion Channel Extracytosolic/Lumenal Domain.
Viet, K.K., Wagner, A., Schwickert, K., Hellwig, N., Brennich, M., Bader, N., Schirmeister, T., Morgner, N., Schindelin, H., Hellmich, U.A.(2019) Structure 27: 1246-1257.e5
- PubMed: 31178222
- DOI: https://doi.org/10.1016/j.str.2019.04.016
- Primary Citation of Related Structures:
6HRR, 6HRS - PubMed Abstract:
TRPML2 is the least structurally characterized mammalian transient receptor potential mucolipin ion channel. The TRPML family hallmark is a large extracytosolic/lumenal domain (ELD) between transmembrane helices S1 and S2. We present crystal structures of the tetrameric human TRPML2 ELD at pH 6.5 (2.0 Å) and 4.5 (2.95 Å), corresponding to the pH values in recycling endosomes and lysosomes. Isothermal titration calorimetry shows Ca 2+ binding to the highly acidic central pre-pore loop which is abrogated at low pH, in line with a pH-dependent channel regulation model. Small angle X-ray scattering confirms the ELD dimensions in solution. Changes in pH or Ca 2+ concentration do not affect the protein's secondary structure, but can influence ELD oligomer integrity according to native mass spectrometry. Our data thus complete the set of high-resolution views of human TRPML channel ELDs and reveal some structural responses to the conditions the TRPML2 ELD encounters as the channel traffics through the endolysosomal system.
Organizational Affiliation:
Institute for Pharmacy and Biochemistry, Johannes Gutenberg-Universität Mainz, 55128 Mainz, Germany; Center for Biomolecular Magnetic Resonance (BMRZ), Goethe-Universität, 60438 Frankfurt am Main, Germany.