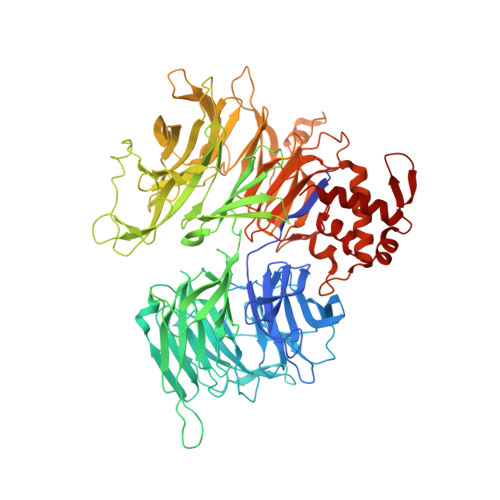
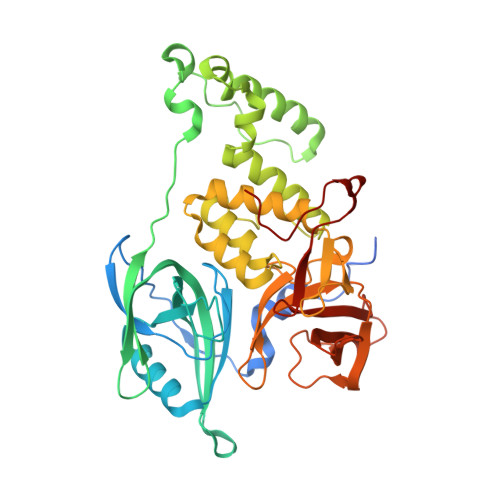
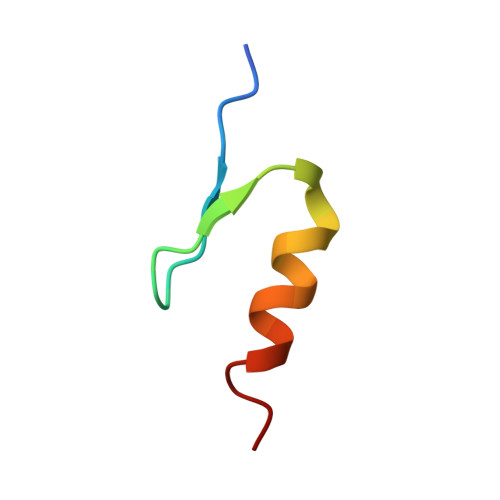
Defining the human C2H2 zinc finger degrome targeted by thalidomide analogs through CRBN.
Sievers, Q.L., Petzold, G., Bunker, R.D., Renneville, A., Slabicki, M., Liddicoat, B.J., Abdulrahman, W., Mikkelsen, T., Ebert, B.L., Thoma, N.H.(2018) Science 362
- PubMed: 30385546
- DOI: https://doi.org/10.1126/science.aat0572
- Primary Citation of Related Structures:
6H0F, 6H0G - PubMed Abstract:
The small molecules thalidomide, lenalidomide, and pomalidomide induce the ubiquitination and proteasomal degradation of the transcription factors Ikaros (IKZF1) and Aiolos (IKZF3) by recruiting a Cys 2 -His 2 (C2H2) zinc finger domain to Cereblon (CRBN), the substrate receptor of the CRL4 CRBN E3 ubiquitin ligase. We screened the human C2H2 zinc finger proteome for degradation in the presence of thalidomide analogs, identifying 11 zinc finger degrons. Structural and functional characterization of the C2H2 zinc finger degrons demonstrates how diverse zinc finger domains bind the permissive drug-CRBN interface. Computational zinc finger docking and biochemical analysis predict that more than 150 zinc fingers bind the drug-CRBN complex in vitro, and we show that selective zinc finger degradation can be achieved through compound modifications. Our results provide a rationale for therapeutically targeting transcription factors that were previously considered undruggable.
Organizational Affiliation:
Broad Institute of Harvard and MIT, Cambridge, MA 02142, USA.