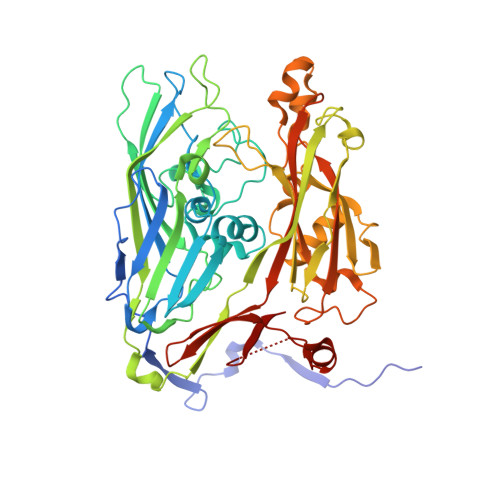
Capsid protein structure, self-assembly, and processing reveal morphogenesis of the marine virophage mavirus.
Born, D., Reuter, L., Mersdorf, U., Mueller, M., Fischer, M.G., Meinhart, A., Reinstein, J.(2018) Proc Natl Acad Sci U S A 115: 7332-7337
- PubMed: 29941605
- DOI: https://doi.org/10.1073/pnas.1805376115
- Primary Citation of Related Structures:
6G41, 6G42, 6G43, 6G44, 6G45 - PubMed Abstract:
Virophages have the unique property of parasitizing giant viruses within unicellular hosts. Little is understood about how they form infectious virions in this tripartite interplay. We provide mechanistic insights into assembly and maturation of mavirus, a marine virophage, by combining structural and stability studies on capsomers, virus-like particles (VLPs), and native virions. We found that the mavirus protease processes the double jelly-roll (DJR) major capsid protein (MCP) at multiple C-terminal sites and that these sites are conserved among virophages. Mavirus MCP assembled in Escherichia coli in the absence and presence of penton protein, forming VLPs with defined size and shape. While quantifying VLPs in E. coli lysates, we found that full-length rather than processed MCP is the competent state for capsid assembly. Full-length MCP was thermally more labile than truncated MCP, and crystal structures of both states indicate that full-length MCP has an expanded DJR core. Thus, we propose that the MCP C-terminal domain serves as a scaffolding domain by adding strain on MCP to confer assembly competence. Mavirus protease processed MCP more efficiently after capsid assembly, which provides a regulation mechanism for timing capsid maturation. By analogy to Sputnik and adenovirus, we propose that MCP processing renders mavirus particles infection competent by loosening interactions between genome and capsid shell and destabilizing pentons for genome release into host cells. The high structural similarity of mavirus and Sputnik capsid proteins together with conservation of protease and MCP processing suggest that assembly and maturation mechanisms described here are universal for virophages.
Organizational Affiliation:
Department of Biomolecular Mechanisms, Max Planck Institute for Medical Research, 69120 Heidelberg, Germany.