Crystal structure of TIA-1 RRM1 in complex with U1C
Jagtap, P.K.A., Sattler, M.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Nucleolysin TIA-1 isoform p40,U1 small nuclear ribonucleoprotein C | 141 | Homo sapiens | Mutation(s): 0 Gene Names: TIA1, SNRPC | 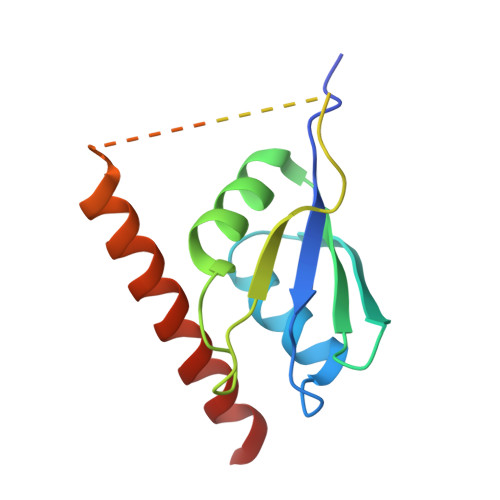 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P09234 (Homo sapiens) Explore P09234 Go to UniProtKB: P09234 | |||||
PHAROS: P09234 GTEx: ENSG00000124562 | |||||
Find proteins for P31483 (Homo sapiens) Explore P31483 Go to UniProtKB: P31483 | |||||
PHAROS: P31483 GTEx: ENSG00000116001 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Groups | P09234P31483 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 71.11 | α = 90 |
| b = 71.11 | β = 90 |
| c = 60.68 | γ = 120 |
| Software Name | Purpose |
|---|---|
| XSCALE | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| XDS | data reduction |
| PHASER | phasing |