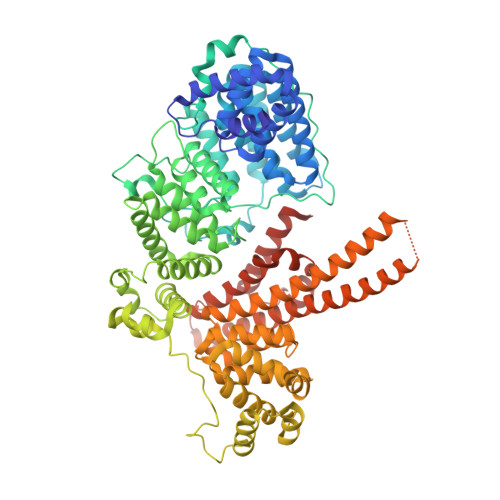
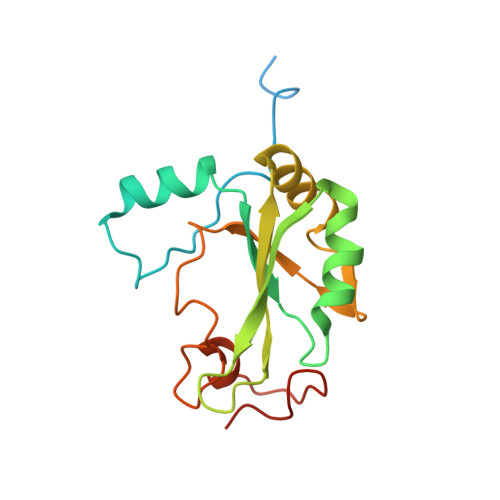
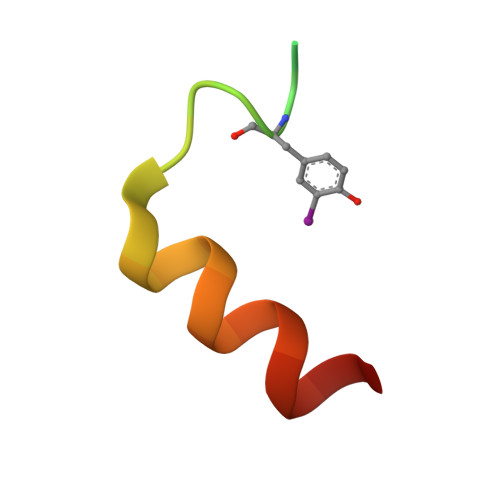
Transcriptional coactivator PGC-1 alpha contains a novel CBP80-binding motif that orchestrates efficient target gene expression.
Cho, H., Rambout, X., Gleghorn, M.L., Nguyen, P.Q.T., Phipps, C.R., Miyoshi, K., Myers, J.R., Kataoka, N., Fasan, R., Maquat, L.E.(2018) Genes Dev 32: 555-567
- PubMed: 29654059
- DOI: https://doi.org/10.1101/gad.309773.117
- Primary Citation of Related Structures:
6D0Y - PubMed Abstract:
Although peroxisome proliferator-activated receptor-γ (PPARγ) coactivator 1α (PGC-1α) is a well-established transcriptional coactivator for the metabolic adaptation of mammalian cells to diverse physiological stresses, the molecular mechanism by which it functions is incompletely understood. Here we used in vitro binding assays, X-ray crystallography, and immunoprecipitations of mouse myoblast cell lysates to define a previously unknown cap-binding protein 80 (CBP80)-binding motif (CBM) in the C terminus of PGC-1α. We show that the CBM, which consists of a nine-amino-acid α helix, is critical for the association of PGC-1α with CBP80 at the 5' cap of target transcripts. Results from RNA sequencing demonstrate that the PGC-1α CBM promotes RNA synthesis from promyogenic genes. Our findings reveal a new conduit between DNA-associated and RNA-associated proteins that functions in a cap-binding protein surveillance mechanism, without which efficient differentiation of myoblasts to myotubes fails to occur.
Organizational Affiliation:
Department of Biochemistry and Biophysics, School of Medicine and Dentistry, University of Rochester, Rochester, New York 14642, USA.