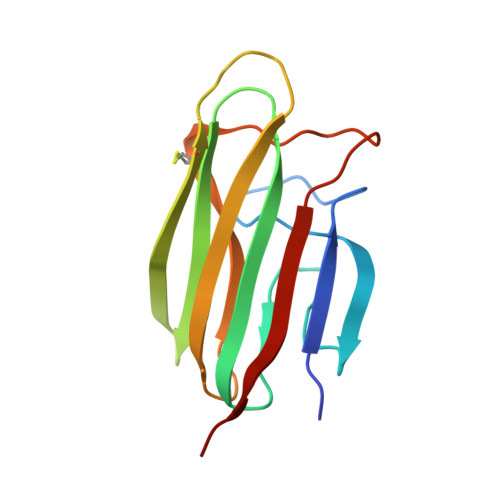
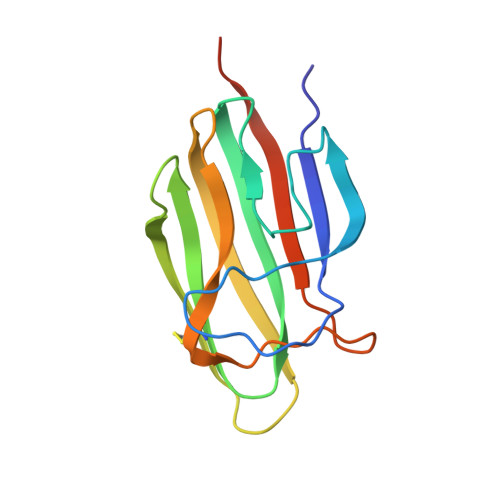
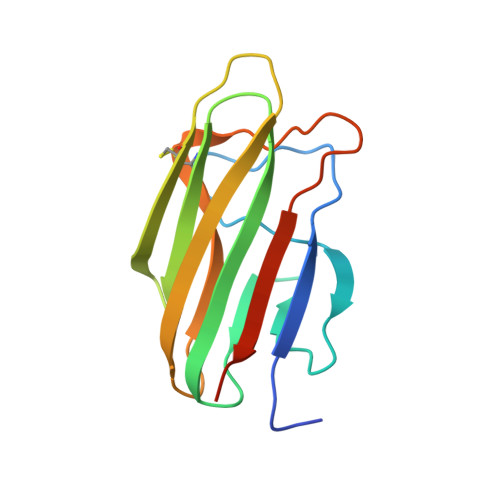
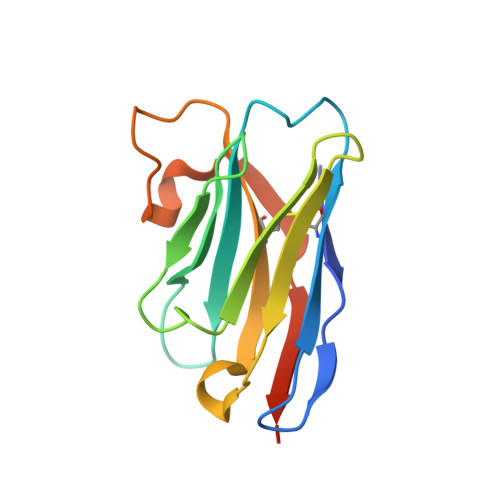
Functional and Structural Characterization of a Potent C1q Inhibitor Targeting the Classical Pathway of the Complement System.
Laursen, N.S., Pedersen, D.V., Gytz, H., Zarantonello, A., Bernth Jensen, J.M., Hansen, A.G., Thiel, S., Andersen, G.R.(2020) Front Immunol 11: 1504-1504
- PubMed: 32849513
- DOI: https://doi.org/10.3389/fimmu.2020.01504
- Primary Citation of Related Structures:
6Z6V - PubMed Abstract:
The classical pathway of complement is important for protection against pathogens and in maintaining tissue homeostasis, but excessive or aberrant activation is directly linked to numerous pathologies. We describe the development and in vitro characterization of C1qNb75, a single domain antibody (nanobody) specific for C1q, the pattern recognition molecule of the classical pathway. C1qNb75 binds to the globular head modules of human C1q with sub-nanomolar affinity and impedes classical pathway mediated hemolysis by IgG and IgM. Crystal structure analysis revealed that C1qNb75 recognizes an epitope primarily located in the C1q B-chain that overlaps with the binding sites of IgG and IgM. Thus, C1qNb75 competitively prevents C1q from binding to IgG and IgM causing blockade of complement activation by the classical pathway. Overall, C1qNb75 represents a high-affinity nanobody-based inhibitor of IgG- and IgM-mediated activation of the classical pathway and may serve as a valuable reagent in mechanistic and functional studies of complement, and as an efficient inhibitor of complement under conditions of excessive CP activation.
Organizational Affiliation:
Department of Molecular Biology and Genetics, Center for Structural Biology, Aarhus University, Aarhus, Denmark.