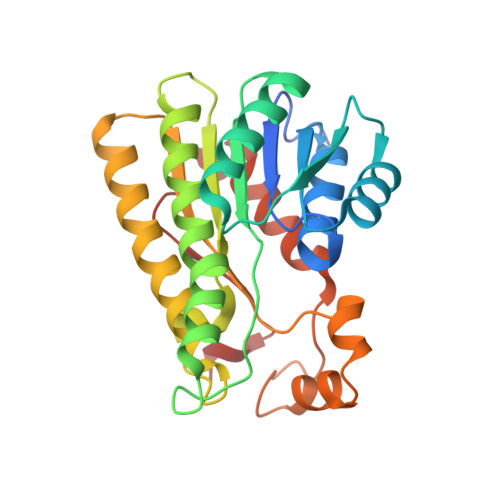
Crystal Contact Engineering Enables Efficient Capture and Purification of an Oxidoreductase by Technical Crystallization.
Grob, P., Huber, M., Walla, B., Hermann, J., Janowski, R., Niessing, D., Hekmat, D., Weuster-Botz, D.(2020) Biotechnol J 15: e2000010-e2000010
- PubMed: 32302461
- DOI: https://doi.org/10.1002/biot.202000010
- Primary Citation of Related Structures:
6Y0S, 6Y15, 6Y1B - PubMed Abstract:
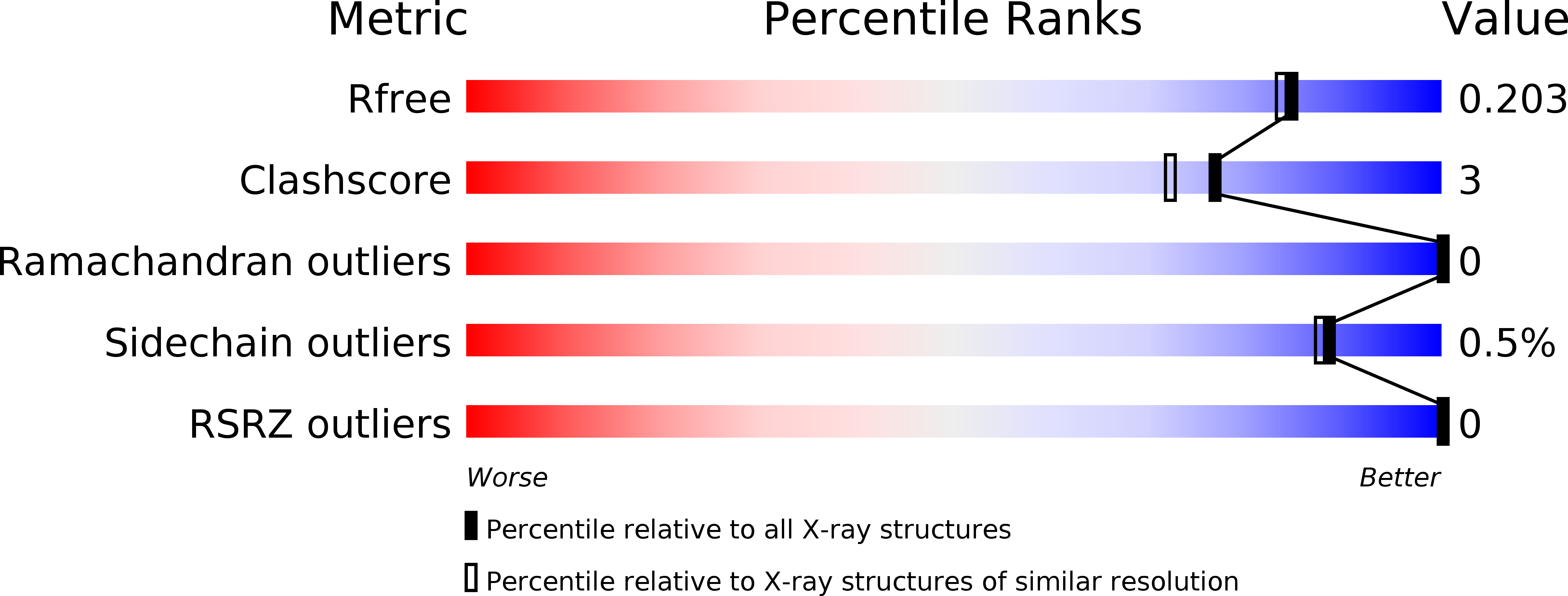
Technical crystallization is an attractive method to purify recombinant proteins. However, it is rarely applied due to the limited crystallizability of many proteins. To overcome this limitation, single amino acid exchanges are rationally introduced to enhance intermolecular interactions at the crystal contacts of the industrially relevant biocatalyst Lactobacillus brevis alcohol dehydrogenase (LbADH). The wildtype (WT) and the best crystallizing and enzymatically active LbADH mutants K32A, D54F, Q126H, and T102E are produced with Escherichia coli and subsequently crystallized from cell lysate in stirred mL-crystallizers. Notwithstanding the high host cell protein (HCP) concentrations in the lysate, all mutants crystallize significantly faster than the WT. Combinations of mutations result in double mutants with faster crystallization kinetics than the respective single mutants, demonstrating a synergetic effect. The almost entire depletion of the soluble LbADH fraction at crystallization equilibrium is observed, proving high yields. The HCP concentration is reduced to below 0.5% after crystal dissolution and recrystallization, and thus a 100-fold HCP reduction is achieved after two successive crystallization steps. The combination of fast kinetics, high yields, and high target protein purity highlights the potential of crystal contact engineering to transform technical crystallization into an efficient protein capture and purification step in biotechnological downstream processes.
Organizational Affiliation:
Technische Universität München, Lehrstuhl für Bioverfahrenstechnik, Boltzmannstraße 15, Garching, 85748, Germany.