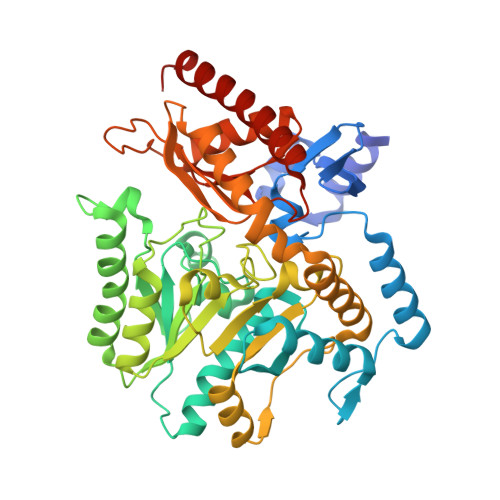
Insight into the dimer dissociation process of the Chromobacterium violaceum (S)-selective amine transaminase.
Ruggieri, F., Campillo-Brocal, J.C., Chen, S., Humble, M.S., Walse, B., Logan, D.T., Berglund, P.(2019) Sci Rep 9: 16946-16946
- PubMed: 31740704
- DOI: https://doi.org/10.1038/s41598-019-53177-3
- Primary Citation of Related Structures:
6S4G - PubMed Abstract:
One of the main factors hampering the implementation in industry of transaminase-based processes for the synthesis of enantiopure amines is their often low storage and operational stability. Our still limited understanding of the inactivation processes undermining the stability of wild-type transaminases represents an obstacle to improving their stability through enzyme engineering. In this paper we present a model describing the inactivation process of the well-characterized (S)-selective amine transaminase from Chromobacterium violaceum. The cornerstone of the model, supported by structural, computational, mutagenesis and biophysical data, is the central role of the catalytic lysine as a conformational switch. Upon breakage of the lysine-PLP Schiff base, the strain associated with the catalytically active lysine conformation is dissipated in a slow relaxation process capable of triggering the known structural rearrangements occurring in the holo-to-apo transition and ultimately promoting dimer dissociation. Due to the occurrence in the literature of similar PLP-dependent inactivation models valid for other non-transaminase enzymes belonging to the same fold-class, the role of the catalytic lysine as conformational switch might extend beyond the transaminase enzyme group and offer new insight to drive future non-trivial engineering strategies.
Organizational Affiliation:
Department of Industrial Biotechnology, KTH Royal Institute of Technology, AlbaNova University Center, SE-106 91, Stockholm, Sweden.