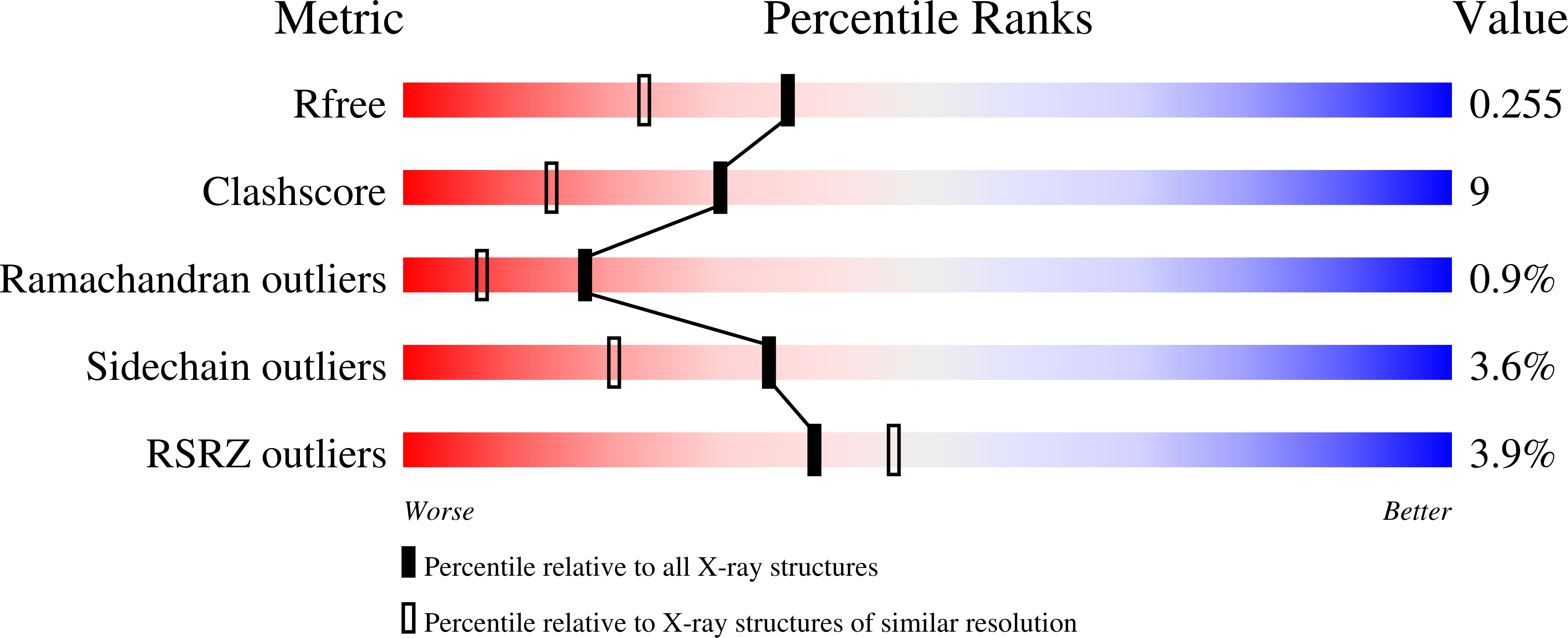
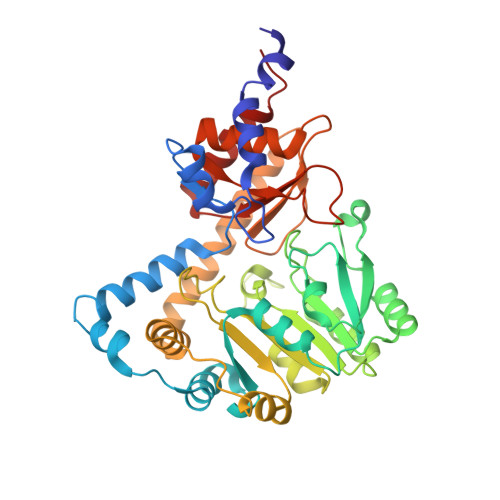
Structural and functional analysis of an l-serine O-phosphate decarboxylase involved in norcobamide biosynthesis.
Keller, S., Wetterhorn, K.M., Vecellio, A., Seeger, M., Rayment, I., Schubert, T.(2019) FEBS Lett 593: 3040-3053
- PubMed: 31325159
- DOI: https://doi.org/10.1002/1873-3468.13543
- Primary Citation of Related Structures:
6OUX - PubMed Abstract:
Structural diversity of natural cobamides (Cbas, B 12 vitamers) is limited to the nucleotide loop. The loop is connected to the cobalt-containing corrin ring via an (R)-1-aminopropan-2-ol O-2-phosphate (AP-P) linker moiety. AP-P is produced by the l-threonine O-3-phosphate (l-Thr-P) decarboxylase CobD. Here, the CobD homolog SMUL_1544 of the organohalide-respiring epsilonproteobacterium Sulfurospirillum multivorans was characterized as a decarboxylase that produces ethanolamine O-phosphate (EA-P) from l-serine O-phosphate (l-Ser-P). EA-P is assumed to serve as precursor of the linker moiety of norcobamides that function as cofactors in the respiratory reductive dehalogenase. SMUL_1544 (SmCobD) is a pyridoxal-5'-phosphate (PLP)-containing enzyme. The structural analysis of the SmCobD apoprotein combined with the characterization of truncated mutant proteins uncovered a role of the SmCobD N-terminus in efficient l-Ser-P conversion.
Organizational Affiliation:
Department of Microbial Interactions, Institute of Microbiology, Friedrich Schiller University, Jena, Germany.