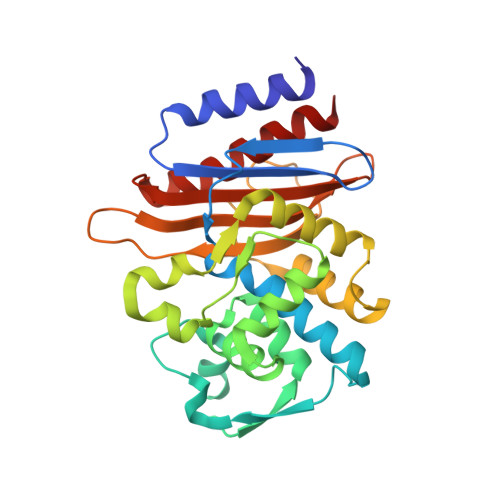
The crystal structure of ESBL TLA-1 in complex with clavulanic acid reveals a second acylation site.
Cifuentes-Castro, V., Rodriguez-Almazan, C., Silva-Sanchez, J., Rudino-Pinera, E.(2020) Biochem Biophys Res Commun 522: 545-551
- PubMed: 31780261
- DOI: https://doi.org/10.1016/j.bbrc.2019.11.138
- Primary Citation of Related Structures:
6NVT, 6NVU, 6PQ8, 6PQ9 - PubMed Abstract:
β-lactamases are the main molecules responsible for giving bacterial resistance against β-lactam antibiotics. The study of β-lactamases has allowed the development of antibiotics capable of inhibiting these enzymes. In this context, extended spectrum β-lactamase (ESBL) TLA-1 has spread in Escherichia coli and Enterobacter cloacae clinical isolates during the last 30 years in Mexico. In this research, the 3D structures of ESBL TLA-1 and TLA-1 S70G mutant, both ligand-free and in complex with clavulanic acid were determined by X-ray crystallography. Four clavulanic acid molecules were found in the structure of TLA-1, two of those were intermediaries of the acylation process and were localized covalently bound to two different amino acid residues, Ser70 and Ser237. The coordinates of TLA-1 in complex with clavulanic acid shows the existence of a second acylation site, additional to Ser70, which might be extendable to several members of the subclass A β-lactamases family. This is the first time that two serines involved in binding clavulanic acid has been reported and described to an atomic level.
Organizational Affiliation:
Departamento de Medicina Molecular y Bioprocesos, Instituto de Biotecnología, Universidad Nacional Autónoma de México, Av. Universidad 2001, C.P. 62210, Cuernavaca, Morelos, Mexico.