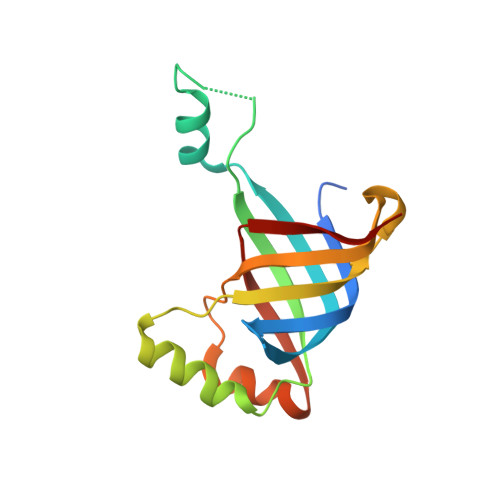
Crystal Structures of R-Type Bacteriocin Sheath and Tube Proteins CD1363 and CD1364 FromClostridium difficilein the Pre-assembled State.
Schwemmlein, N., Pippel, J., Gazdag, E.M., Blankenfeldt, W.(2018) Front Microbiol 9: 1750-1750
- PubMed: 30127773
- DOI: https://doi.org/10.3389/fmicb.2018.01750
- Primary Citation of Related Structures:
6GKW, 6GKX - PubMed Abstract:
Diffocins are high-molecular-weight phage tail-like bacteriocins (PTLBs) that some Clostridium difficile strains produce in response to SOS induction. Similar to the related R-type pyocins from Pseudomonas aeruginosa , R-type diffocins act as molecular puncture devices that specifically penetrate the cell envelope of other C. difficile strains to dissipate the membrane potential and kill the attacked bacterium. Thus, R-type diffocins constitute potential therapeutic agents to counter C. difficile -associated infections. PTLBs consist of rigid and contractile protein complexes. They are composed of a baseplate, receptor-binding tail fibers and an inner needle-like tube surrounded by a contractile sheath. In the mature particle, the sheath and tube structure form a complex network comprising up to 200 copies of a sheath and a tube protein each. Here, we report the crystal structures together with small angle X-ray scattering data of the sheath and tube proteins CD1363 (39 kDa) and CD1364 (16 kDa) from C. difficile strain CD630 in a monomeric pre-assembly form at 1.9 and 1.5 Å resolution, respectively. The tube protein CD1364 displays a compact fold and shares highest structural similarity with a tube protein from Bacillus subtilis but is remarkably different from that of the R-type pyocin from P. aeruginosa . The structure of the R-type diffocin sheath protein, on the other hand, is highly conserved. It contains two domains, whereas related members such as bacteriophage tail sheath proteins comprise up to four, indicating that R-type PTLBs may represent the minimal protein required for formation of a complete sheath structure. Comparison of CD1363 and CD1364 with structures of PTLBs and related assemblies suggests that several conformational changes are required to form complete assemblies. In the sheath, rearrangement of the flexible N- and C-terminus enables extensive interactions between the other subunits, whereas for the tube, such contacts are primarily established by mobile α-helices. Together, our results combined with information from structures of homologous assemblies allow constructing a preliminary model of the sheath and tube assembly from R-type diffocin.
Organizational Affiliation:
Structure and Function of Proteins, Helmholtz Centre for Infection Research, Braunschweig, Germany.