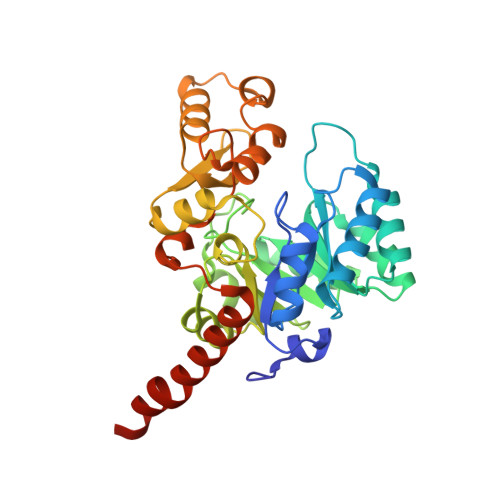
Steric hindrance controls pyridine nucleotide specificity of a flavin-dependent NADH:quinone oxidoreductase.
Ball, J., Reis, R.A.G., Agniswamy, J., Weber, I.T., Gadda, G.(2019) Protein Sci 28: 167-175
- PubMed: 30246917
- DOI: https://doi.org/10.1002/pro.3514
- Primary Citation of Related Structures:
6E2A - PubMed Abstract:
The crystal structure of the NADH:quinone oxidoreductase PA1024 has been solved in complex with NAD + to 2.2 Å resolution. The nicotinamide C4 is 3.6 Å from the FMN N5 atom, with a suitable orientation for facile hydride transfer. NAD + binds in a folded conformation at the interface of the TIM-barrel domain and the extended domain of the enzyme. Comparison of the enzyme-NAD + structure with that of the ligand-free enzyme revealed a different conformation of a short loop (75-86) that is part of the NAD + -binding pocket. P78, P82, and P84 provide internal rigidity to the loop, whereas Q80 serves as an active site latch that secures the NAD + within the binding pocket. An interrupted helix consisting of two α-helices connected by a small three-residue loop binds the pyrophosphate moiety of NAD + . The adenine moiety of NAD + appears to π-π stack with Y261. Steric constraints between the adenosine ribose of NAD + , P78, and Q80, control the strict specificity of the enzyme for NADH. Charged residues do not play a role in the specificity of PA1024 for the NADH substrate.
Organizational Affiliation:
Department of Chemistry, Georgia State University, Atlanta, Georgia, 30302-3965.