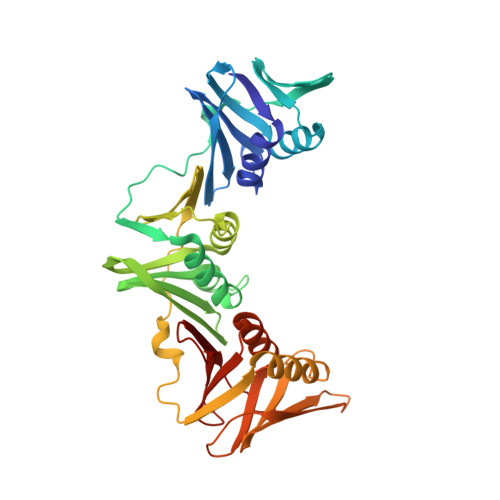
Crystal structures and biochemical characterization of DNA sliding clamps from three Gram-negative bacterial pathogens.
McGrath, A.E., Martyn, A.P., Whittell, L.R., Dawes, F.E., Beck, J.L., Dixon, N.E., Kelso, M.J., Oakley, A.J.(2018) J Struct Biol 204: 396-405
- PubMed: 30366028
- DOI: https://doi.org/10.1016/j.jsb.2018.10.008
- Primary Citation of Related Structures:
6AMQ, 6AMS, 6AP4 - PubMed Abstract:
Bacterial sliding clamps bind to DNA and act as protein-protein interaction hubs for several proteins involved in DNA replication and repair. The partner proteins all bind to a common pocket on sliding clamps via conserved linear peptide sequence motifs, which suggest the pocket as an attractive target for development of new antibiotics. Herein we report the X-ray crystal structures and biochemical characterization of β sliding clamps from the Gram-negative pathogens Pseudomonas aeruginosa, Acinetobacter baumannii and Enterobacter cloacae. The structures reveal close similarity between the pathogen and Escherichia coli clamps and similar patterns of binding to linear clamp-binding motif peptides. The results suggest that linear motif-sliding clamp interactions are well conserved and an antibiotic targeting the sliding clamp should have broad-spectrum activity against Gram-negative pathogens.
Organizational Affiliation:
Molecular Horizons and School of Chemistry and Molecular Bioscience, University of Wollongong, and Illawarra Health and Medical Research Institute, Wollongong, New South Wales, Australia.