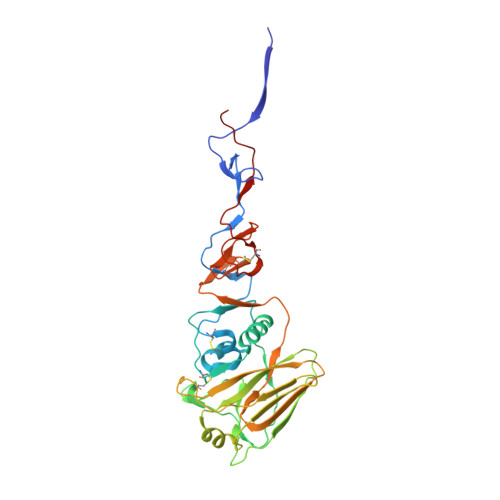
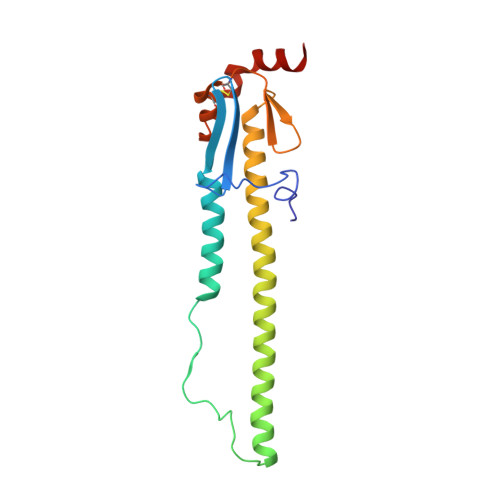
Structural basis of influenza virus fusion inhibition by the antiviral drug Arbidol.
Kadam, R.U., Wilson, I.A.(2017) Proc Natl Acad Sci U S A 114: 206-214
- PubMed: 28003465
- DOI: https://doi.org/10.1073/pnas.1617020114
- Primary Citation of Related Structures:
5T6N, 5T6S - PubMed Abstract:
The broad-spectrum antiviral drug Arbidol shows efficacy against influenza viruses by targeting the hemagglutinin (HA) fusion machinery. However, the structural basis of the mechanism underlying fusion inhibition by Arbidol has remained obscure, thereby hindering its further development as a specific and optimized influenza therapeutic. We determined crystal structures of Arbidol in complex with influenza virus HA from pandemic 1968 H3N2 and recent 2013 H7N9 viruses. Arbidol binds in a hydrophobic cavity in the HA trimer stem at the interface between two protomers. This cavity is distal to the conserved epitope targeted by broadly neutralizing stem antibodies and is ∼16 Å from the fusion peptide. Arbidol primarily makes hydrophobic interactions with the binding site but also induces some conformational rearrangements to form a network of inter- and intraprotomer salt bridges. By functioning as molecular glue, Arbidol stabilizes the prefusion conformation of HA that inhibits the large conformational rearrangements associated with membrane fusion in the low pH of the endosome. This unique binding mode compared with the small-molecule inhibitors of other class I fusion proteins enhances our understanding of how small molecules can function as fusion inhibitors and guides the development of broad-spectrum therapeutics against influenza virus.
Organizational Affiliation:
Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037.