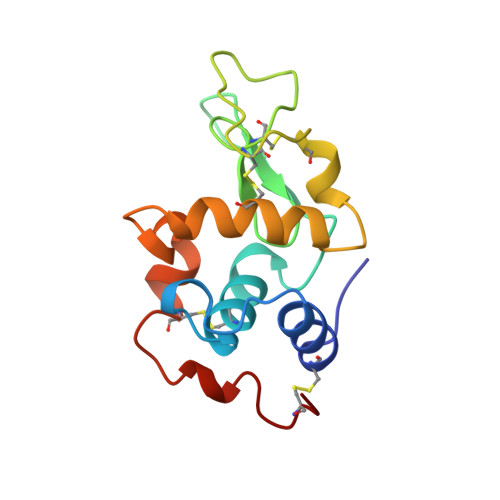
A Rare Lysozyme Crystal Form Solved Using Highly Redundant Multiple Electron Diffraction Datasets from Micron-Sized Crystals.
Xu, H., Lebrette, H., Yang, T., Srinivas, V., Hovmoller, S., Hogbom, M., Zou, X.(2018) Structure 26: 667-675.e3
- PubMed: 29551291
- DOI: https://doi.org/10.1016/j.str.2018.02.015
- Primary Citation of Related Structures:
5OCV - PubMed Abstract:
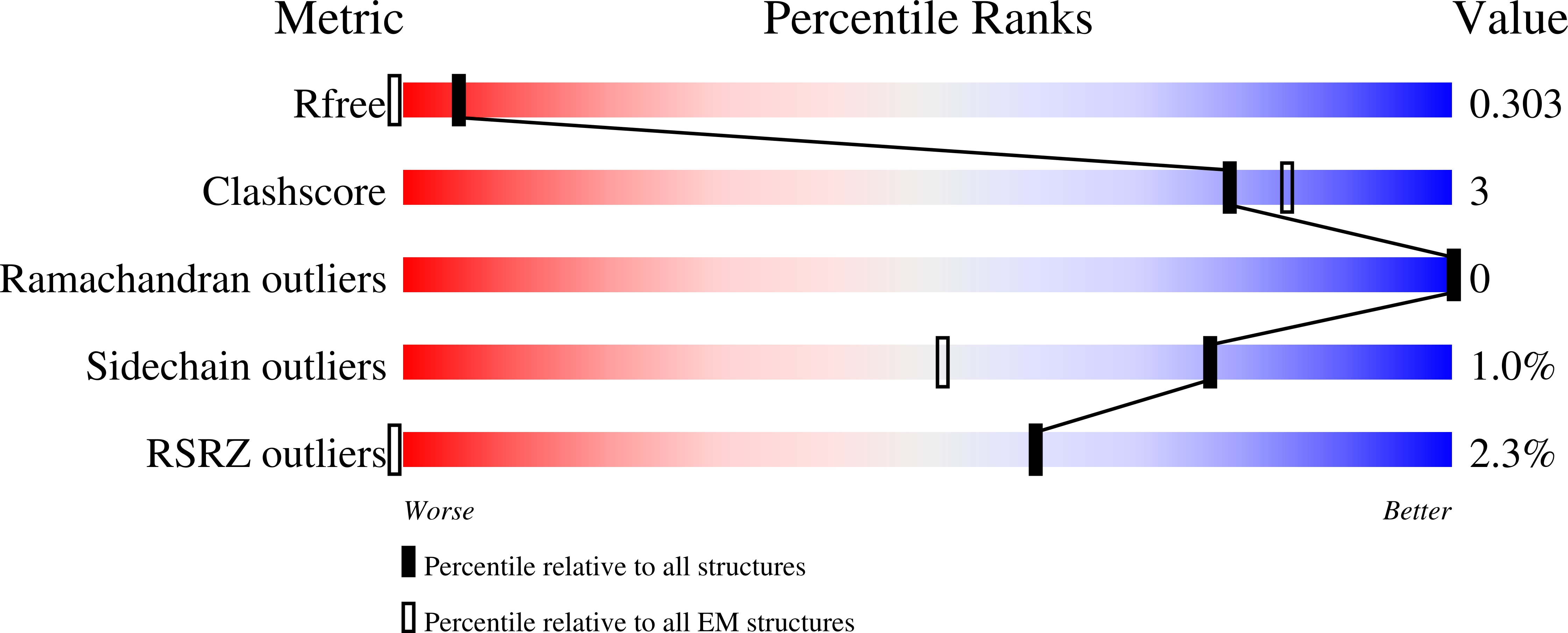
Recent developments of novel electron diffraction techniques have shown to be powerful for determination of atomic resolution structures from micron- and nano-sized crystals, too small to be studied by single-crystal X-ray diffraction. In this work, the structure of a rare lysozyme polymorph is solved and refined using continuous rotation MicroED data and standard X-ray crystallographic software. Data collection was performed on a standard 200 kV transmission electron microscope (TEM) using a highly sensitive detector with a short readout time. The data collection is fast (∼3 min per crystal), allowing multiple datasets to be rapidly collected from a large number of crystals. We show that merging data from 33 crystals significantly improves not only the data completeness, overall I/σ and the data redundancy, but also the quality of the final atomic model. This is extremely useful for electron beam-sensitive crystals of low symmetry or with a preferred orientation on the TEM grid.
Organizational Affiliation:
Inorganic and Structural Chemistry, Department of Materials and Environmental Chemistry, Stockholm University, 106 91 Stockholm, Sweden.