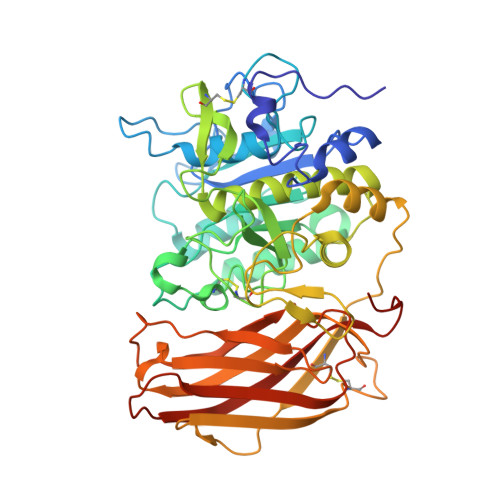
Structure of the unliganded form of the proprotein convertase furin suggests activation by a substrate-induced mechanism.
Dahms, S.O., Arciniega, M., Steinmetzer, T., Huber, R., Than, M.E.(2016) Proc Natl Acad Sci U S A 113: 11196-11201
- PubMed: 27647913
- DOI: https://doi.org/10.1073/pnas.1613630113
- Primary Citation of Related Structures:
5JXG, 5JXH, 5JXI, 5JXJ - PubMed Abstract:
Proprotein convertases (PCs) are highly specific proteases required for the proteolytic modification of many secreted proteins. An unbalanced activity of these enzymes is connected to pathologies like cancer, atherosclerosis, hypercholesterolaemia, and infectious diseases. Novel protein crystallographic structures of the prototypical PC family member furin in different functional states were determined to 1.8-2.0 Å. These, together with biochemical data and modeling by molecular dynamics calculations, suggest essential elements underlying its unusually high substrate specificity. Furin shows a complex activation mechanism and exists in at least four defined states: (i) the "off state," incompatible with substrate binding as seen in the unliganded enzyme; (ii) the active "on state" seen in inhibitor-bound furin; and the respective (iii) calcium-free and (iv) calcium-bound forms. The transition from the off to the on state is triggered by ligand binding at subsites S1 to S4 and appears to underlie the preferential recognition of the four-residue sequence motif of furin. The molecular dynamics simulations of the four structural states reflect the experimental observations in general and provide approximations of the respective stabilities. Ligation by calcium at the PC-specific binding site II influences the active-site geometry and determines the rotamer state of the oxyanion hole-forming Asn295, and thus adds a second level of the activity modulation of furin. The described crystal forms and the observations of different defined functional states may foster the development of new tools and strategies for pharmacological intervention targeting furin.
Organizational Affiliation:
Protein Crystallography Group, Leibniz Institute on Aging-Fritz Lipmann Institute (FLI), 07745 Jena, Germany.