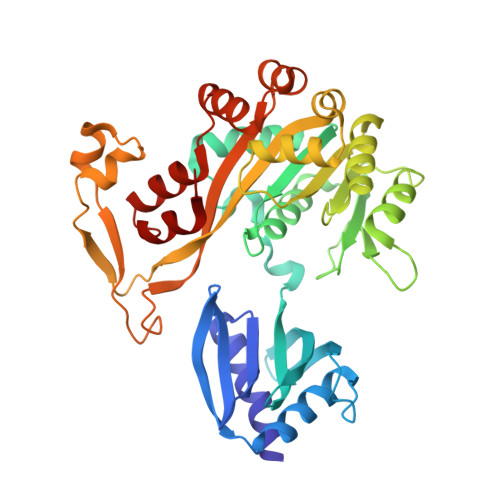
Crystal Structure of a Type IV Pilus Assembly ATPase: Insights into the Molecular Mechanism of PilB from Thermus thermophilus.
Mancl, J.M., Black, W.P., Robinson, H., Yang, Z., Schubot, F.D.(2016) Structure 24: 1886-1897
- PubMed: 27667690
- DOI: https://doi.org/10.1016/j.str.2016.08.010
- Primary Citation of Related Structures:
5IT5 - PubMed Abstract:
Type IV pili (T4P) mediate bacterial motility and virulence. The PilB/GspE family ATPases power the assembly of T4P and type 2 secretion systems. We determined the structure of the ATPase region of PilB (PilB ATP ) in complex with ATPγS to provide a model of a T4P assembly ATPase and a view of a PilB/GspE family hexamer at better than 3-Å resolution. Spatial positioning and conformations of the protomers suggest a mechanism of force generation. All six PilB ATP protomers contain bound ATPγS. Two protomers form a closed conformation poised for ATP hydrolysis. The other four molecules assume an open conformation but separate into two pairs with distinct active-site accessibilities. We propose that one pair represents the post-hydrolysis phase while the other pair appears poised for ADP/ATP exchange. Collectively, the data suggest that T4P assembly is powered by coordinating concurrent substrate binding with ATP hydrolysis across the PilB hexamer.
Organizational Affiliation:
Department of Biological Sciences, Virginia Polytechnic Institute and State University, 125 Life Sciences 1 (MC 0910), 970 Washington Street Southwest, Blacksburg, VA 24061, USA.