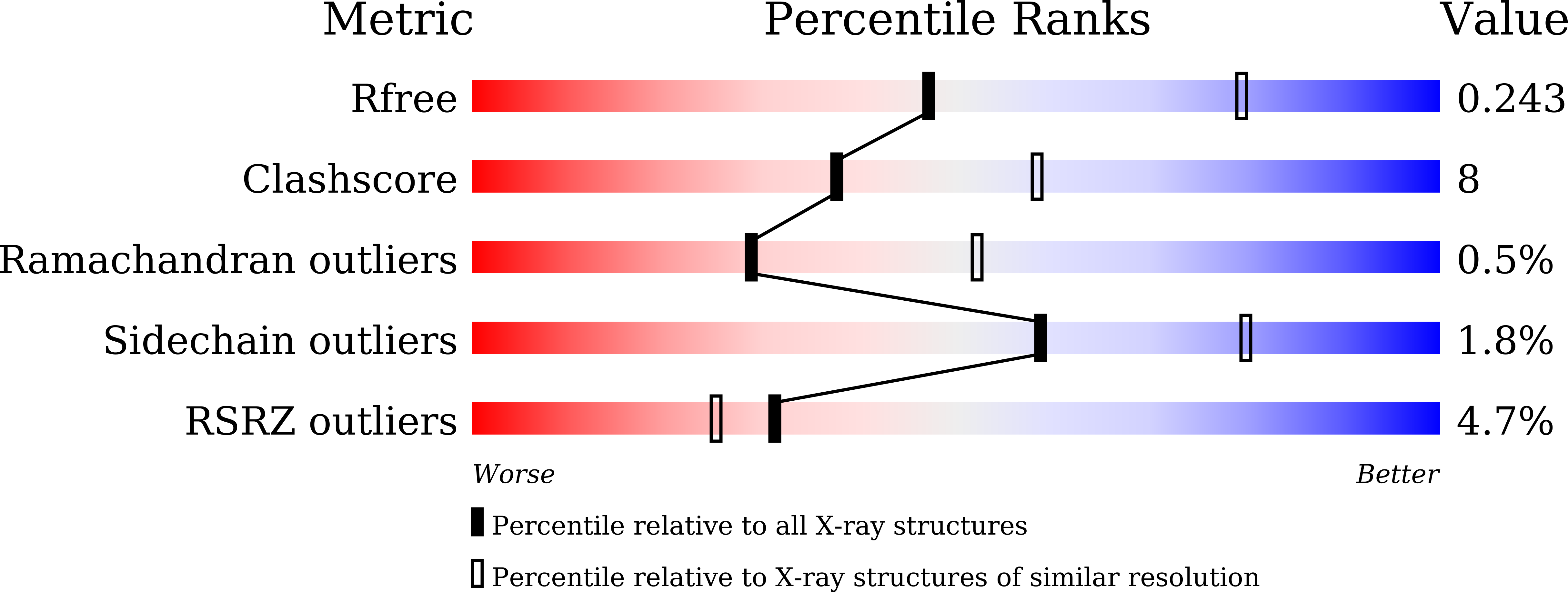
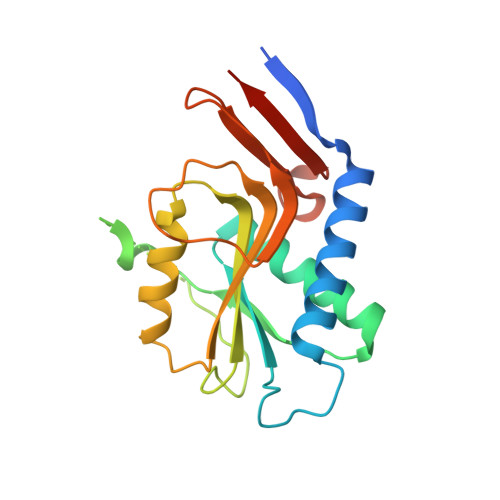
Structure and mapping of spontaneous mutational sites of PyrR from Mycobacterium tuberculosis
Ghode, P., Ramachandran, S., Bifani, P., Sivaraman, J.(2016) Biochem Biophys Res Commun 471: 409-415
- PubMed: 26902118
- DOI: https://doi.org/10.1016/j.bbrc.2016.02.071
- Primary Citation of Related Structures:
5IAO - PubMed Abstract:
The emergence of resistant Mycobacterium tuberculosis (Mtb) infection and the dearth of drugs against tuberculosis have made it imperative to identify and validate novel targets and classes of drugs for treatment. The pyrimidine operon regulatory protein (PyrR), a regulator of de novo pyrimidine synthesis, is an essential enzyme and a probable 5-fluorouracil (5-FU) target in Mtb, with mutations in PyrR attributable to 5-FU resistance. Here we report, for the first time, the co-crystal structure of the PyrR-5-FU complex along with mapping of spontaneous mutational sites of PyrR. A cluster of mutations in the presence of the drug usually indicates a plausible region of drug-target interaction. Notably, we observed that three of the mutated PyrR residues lie in close proximity to the 5-FU binding site, including the amino acid Val178, which is involved in water mediated hydrogen bonding contact with 5-FU. Computational modeling of the PyrR-5'-phosphoribosyl-α-1'-pyrophosphate (PRPP) complex revealed the location of several other mutations at the PRPP binding site of PyrR, indicating their probable role in resistance. Indeed, 5-FU-resistant strains harboring these mutations exhibited decreased susceptibility to 5-FU. Considering that pyrimidine analogs are predominantly regarded to inhibit PyrR, the present studies will be beneficial for the screening of appropriate inhibitors of PyrR and help provide insight into future TB drug design and development.
Organizational Affiliation:
Department of Biological Sciences, National University of Singapore, 14 Science Drive 4, Singapore, 117543, Singapore; Novartis Institute for Tropical Diseases, 10 Biopolis Road, Singapore, 138670, Singapore.