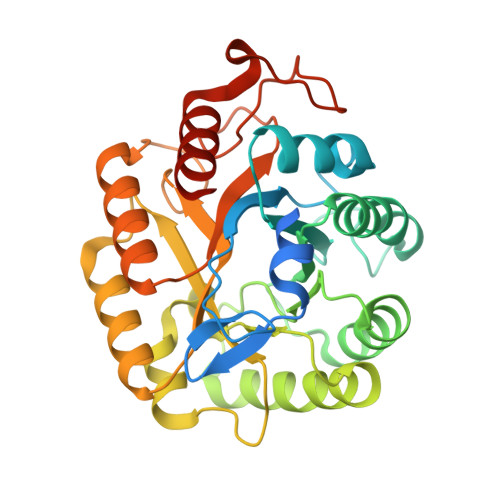
Biochemical and structural characterization of a novel halotolerant cellulase from soil metagenome
Garg, R., Srivastava, R., Brahma, V., Verma, L., Karthikeyan, S., Sahni, G.(2016) Sci Rep 6: 39634-39634
- PubMed: 28008971
- DOI: https://doi.org/10.1038/srep39634
- Primary Citation of Related Structures:
5I2U - PubMed Abstract:
Cellulase catalyzes the hydrolysis of β-1,4-linkages of cellulose to produce industrially relevant monomeric subunits. Cellulases find their applications in pulp and paper, laundry, food and feed, textile, brewing industry and in biofuel production. These industries always have great demand for cellulases that can work efficiently even in harsh conditions such as high salt, heat, and acidic environments. While, cellulases with high thermal and acidic stability are already in use, existence of a high halotolerant cellulase is still elusive. Here, we report a novel cellulase Cel5R, obtained from soil metagenome that shows high halotolerance and thermal stability. The biochemical and functional characterization of Cel5R revealed its endoglucanase activity and high halostability. In addition, the crystal structure of Cel5R determined at 2.2 Å resolution reveals a large number of acidic residues on the surface of the protein that contribute to the halophilic nature of this enzyme. Moreover, we demonstrate that the four free and non-conserved cysteine residues (C65, C90, C231 and C273) contributes to the thermal stability of Cel5R by alanine scanning experiments. Thus, the newly identified endoglucanase Cel5R is a promising candidate for various industrial applications.
Organizational Affiliation:
CSIR-Institute Of Microbial Technology, Council Of Scientific and Industrial Research (CSIR), Sector 39 A, Chandigarh 160036, India.