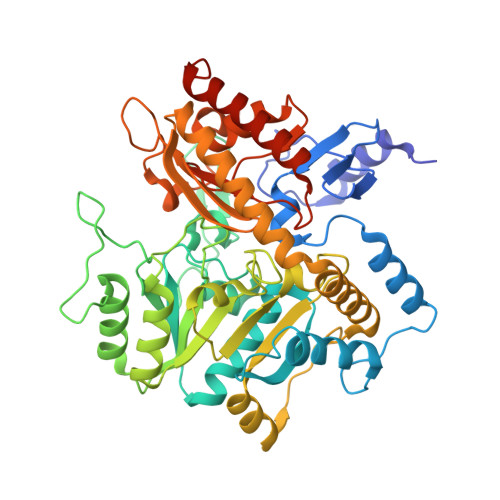
Structural Basis of Substrate Range and Enantioselectivity of Two S-Selective Omega- Transaminases
Van Oosterwijk, N., Willies, S., Hekelaar, J., Terwisscha Van Scheltinga, A.C., Turner, N.J., Dijkstra, B.W.(2016) Biochemistry 55: 4422
- PubMed: 27428867
- DOI: https://doi.org/10.1021/acs.biochem.6b00370
- Primary Citation of Related Structures:
5G09, 5G0A, 5G2P, 5G2Q - PubMed Abstract:
ω-Transaminases are enzymes that can introduce an amino group in industrially interesting compounds. We determined crystal structures of two (S)-selective ω-transaminases, one from Arthrobacter sp. (Ars-ωTA) and one from Bacillus megaterium (BM-ωTA), which have 95% identical sequences but somewhat different activity profiles. Substrate profiling measurements using a range of (R)- and (S)-substrates showed that both enzymes have a preference for substrates with large, flat cyclic side groups, for which the activity of BM-ωTA is generally somewhat higher. BM-ωTA has a preference for (S)-3,3-dimethyl-2-butylamine significantly stronger than that of Ars-ωTA, as well as a weaker enantiopreference for 1-cyclopropylethylamine. The crystal structures showed that, as expected for (S)-selective transaminases, both enzymes have the typical transaminase type I fold and have spacious active sites to accommodate largish substrates. A structure of BM-ωTA with bound (R)-α-methylbenzylamine explains the enzymes' preference for (S)-substrates. Site-directed mutagenesis experiments revealed that the presence of a tyrosine, instead of a cysteine, at position 60 increases the relative activities on several small substrates. A structure of Ars-ωTA with bound l-Ala revealed that the Arg442 side chain has been repositioned to bind the l-Ala carboxylate. Compared to the arginine switch residue in other transaminases, Arg442 is shifted by six residues in the amino acid sequence, which appears to be a consequence of extra loops near the active site that narrow the entrance to the active site.
Organizational Affiliation:
Laboratory of Biophysical Chemistry, Groningen Biomolecular Sciences and Biotechnology Institute (GBB), University of Groningen , Nijenborgh 7, 9747 AG Groningen, The Netherlands.