The Functional Unit of Neisseria meningitidis 3-Deoxy--Arabino-Heptulosonate 7-Phosphate Synthase Is Dimeric.
Cross, P.J., Heyes, L.C., Zhang, S., Nazmi, A.R., Parker, E.J.(2016) PLoS One 11: e0145187-e0145187
- PubMed: 26828675
- DOI: https://doi.org/10.1371/journal.pone.0145187
- Primary Citation of Related Structures:
4UCG, 5DCE - PubMed Abstract:
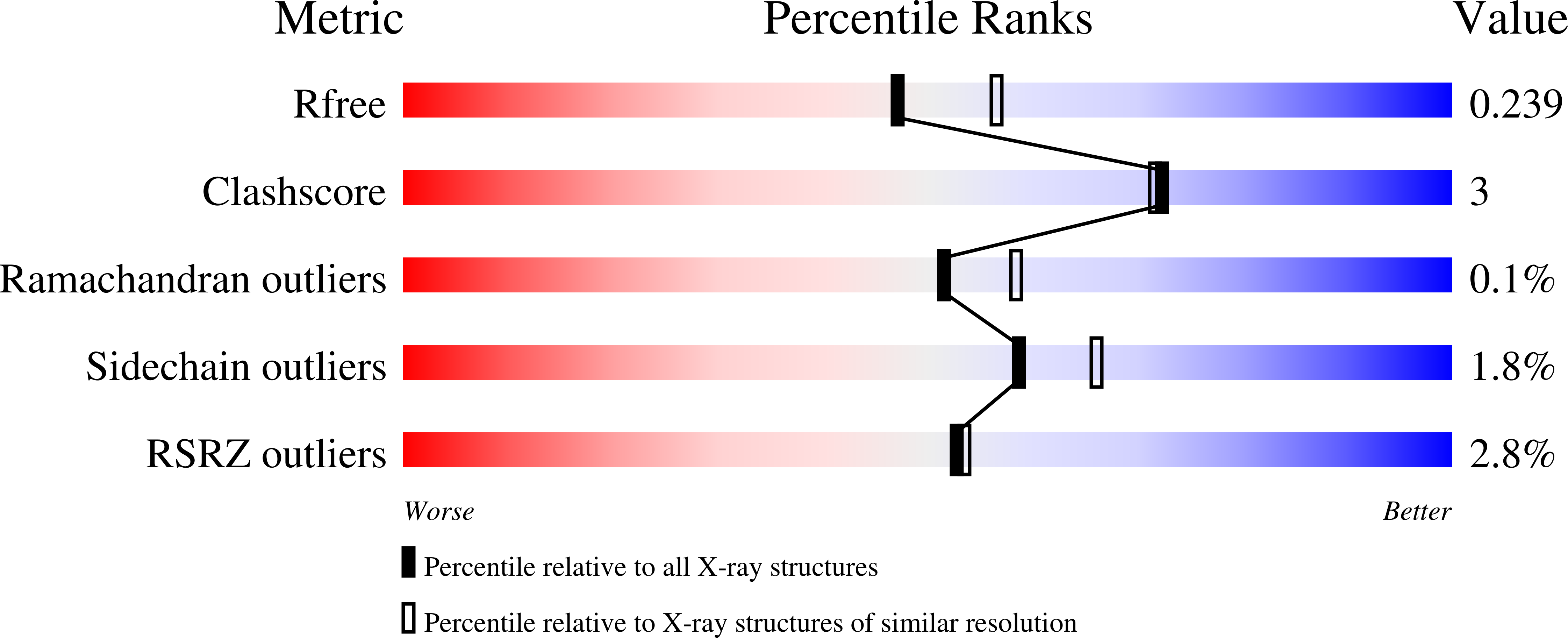
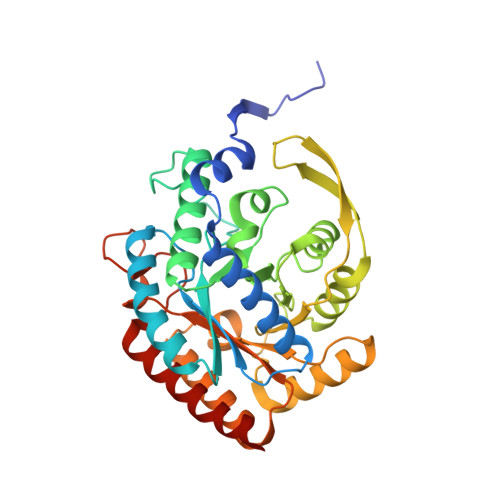
Neisseria meningitidis 3-deoxy-D-arabino-heptulosonate 7-phosphate synthase (NmeDAH7PS) adopts a homotetrameric structure consisting of an extensive and a less extensive interface. Perturbation of the less extensive interface through a single mutation of a salt bridge (Arg126-Glu27) formed at the tetramer interface of all chains resulted in a dimeric DAH7PS in solution, as determined by small angle X-ray scattering, analytical ultracentrifugation and analytical size-exclusion chromatography. The dimeric NmeDAH7PSR126S variant was shown to be catalytically active in the aldol-like condensation reaction between D-erythrose 4-phosphate and phosphoenolpyruvate, and allosterically inhibited by L-phenylalanine to the same extent as the wild-type enzyme. The dimeric NmeDAH7PSR126S variant exhibited a slight reduction in thermal stability by differential scanning calorimetry experiments and a slow loss of activity over time compared to the wild-type enzyme. Although NmeDAH7PSR126S crystallised as a tetramer, like the wild-type enzyme, structural asymmetry at the less extensive interface was observed consistent with its destabilisation. The tetrameric association enabled by this Arg126-Glu27 salt-bridge appears to contribute solely to the stability of the protein, ultimately revealing that the functional unit of NmeDAH7PS is dimeric.
Organizational Affiliation:
Biomolecular Interaction Centre and Department of Chemistry, University of Canterbury, Christchurch, New Zealand.