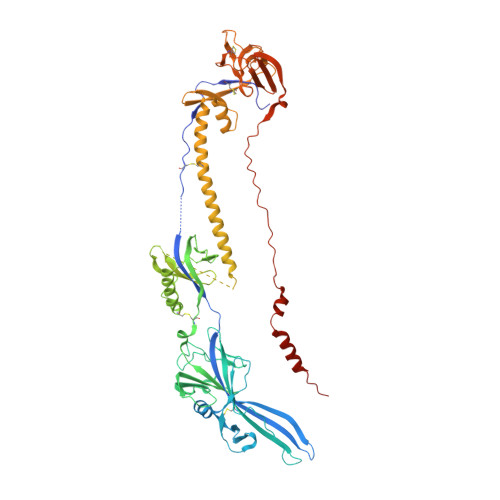
Crystal Structure of the Human Cytomegalovirus Glycoprotein B.
Burke, H.G., Heldwein, E.E.(2015) PLoS Pathog 11: e1005227-e1005227
- PubMed: 26484870
- DOI: https://doi.org/10.1371/journal.ppat.1005227
- Primary Citation of Related Structures:
5CXF - PubMed Abstract:
Human cytomegalovirus (HCMV), a dsDNA, enveloped virus, is a ubiquitous pathogen that establishes lifelong latent infections and caused disease in persons with compromised immune systems, e.g., organ transplant recipients or AIDS patients. HCMV is also a leading cause of congenital viral infections in newborns. Entry of HCMV into cells requires the conserved glycoprotein B (gB), thought to function as a fusogen and reported to bind signaling receptors. gB also elicits a strong immune response in humans and induces the production of neutralizing antibodies although most anti-gB Abs are non-neutralizing. Here, we report the crystal structure of the HCMV gB ectodomain determined to 3.6-Å resolution, which is the first atomic-level structure of any betaherpesvirus glycoprotein. The structure of HCMV gB resembles the postfusion structures of HSV-1 and EBV homologs, establishing it as a new member of the class III viral fusogens. Despite structural similarities, each gB has a unique domain arrangement, demonstrating structural plasticity of gB that may accommodate virus-specific functional requirements. The structure illustrates how extensive glycosylation of the gB ectodomain influences antibody recognition. Antigenic sites that elicit neutralizing antibodies are more heavily glycosylated than those that elicit non-neutralizing antibodies, which suggest that HCMV gB uses glycans to shield neutralizing epitopes while exposing non-neutralizing epitopes. This glycosylation pattern may have evolved to direct the immune response towards generation of non-neutralizing antibodies thus helping HCMV to avoid clearance. HCMV gB structure provides a starting point for elucidation of its antigenic and immunogenic properties and aid in the design of recombinant vaccines and monoclonal antibody therapies.
Organizational Affiliation:
Department of Molecular Biology and Microbiology and Graduate Program in Molecular Microbiology, Sackler School of Graduate Biomedical Sciences, Tufts University School of Medicine, Boston, Massachusetts, United States of America.