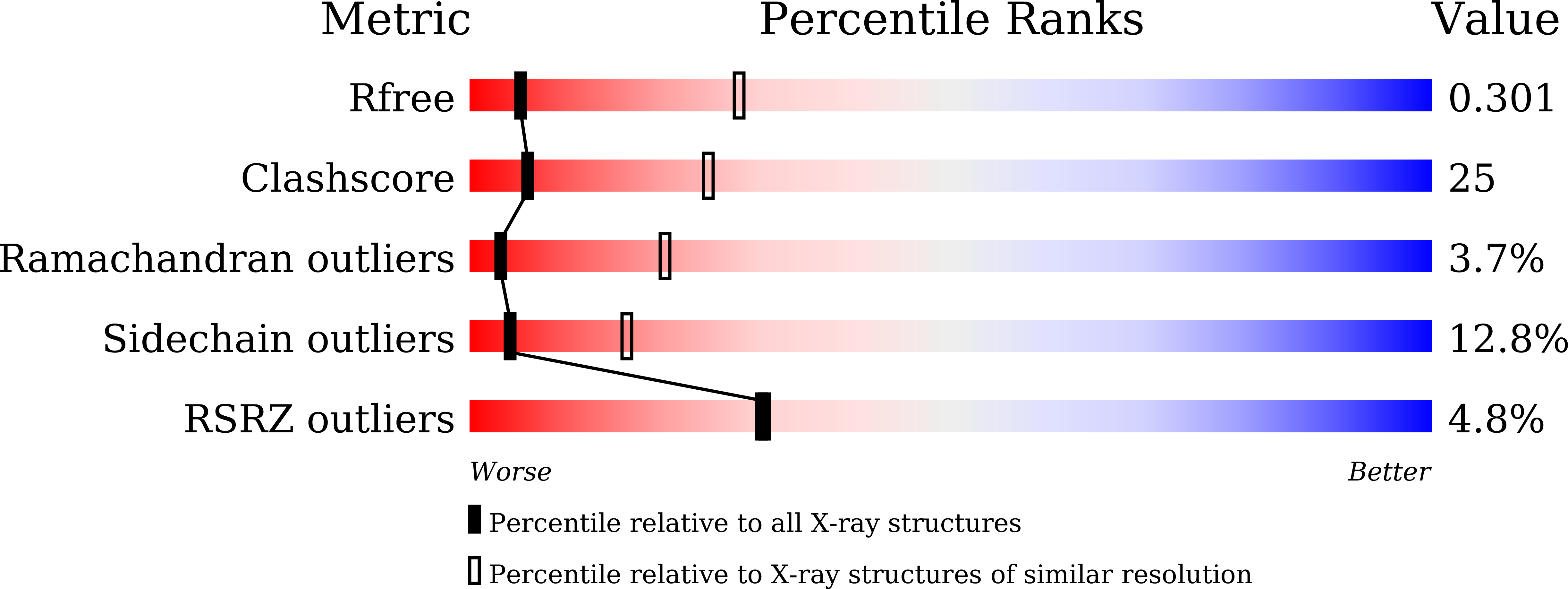
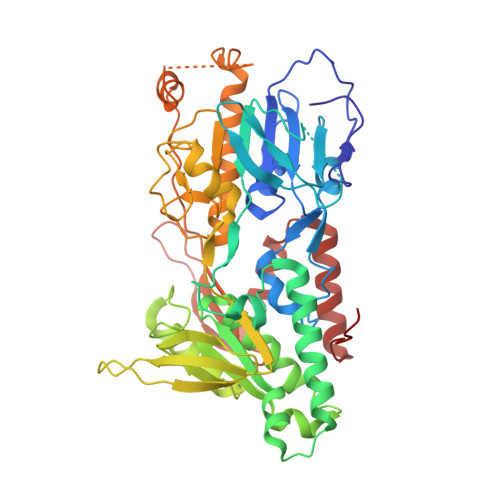
Ubiquinone binding site of yeast NADH dehydrogenase revealed by structures binding novel competitive- and mixed-type inhibitors
Yamashita, T., Inaoka, D.K., Shiba, T., Oohashi, T., Iwata, S., Yagi, T., Kosaka, H., Miyoshi, H., Harada, S., Kita, K., Hirano, K.(2018) Sci Rep 8: 2427-2427
- PubMed: 29402945
- DOI: https://doi.org/10.1038/s41598-018-20775-6
- Primary Citation of Related Structures:
5YJW, 5YJX, 5YJY - PubMed Abstract:
Yeast Ndi1 is a monotopic alternative NADH dehydrogenase. Its crystal structure in complex with the electron acceptor, ubiquinone, has been determined. However, there has been controversy regarding the ubiquinone binding site. To address these points, we identified the first competitive inhibitor of Ndi1, stigmatellin, along with new mixed-type inhibitors, AC0-12 and myxothiazol, and thereby determined the crystal structures of Ndi1 in complexes with the inhibitors. Two separate binding sites of stigmatellin, STG-1 and STG-2, were observed. The electron density at STG-1, located at the vicinity of the FAD cofactor, further demonstrated two binding modes: STG-1a and STG-1b. AC0-12 and myxothiazol are also located at the vicinity of FAD. The comparison of the binding modes among stigmatellin at STG-1, AC0-12, and myxothiazol revealed a unique position for the aliphatic tail of stigmatellin at STG-1a. Mutations of amino acid residues that interact with this aliphatic tail at STG-1a reduced the affinity of Ndi1 for ubiquinone. In conclusion, the position of the aliphatic tail of stigmatellin at STG-1a provides a structural basis for its competitive inhibition of Ndi1. The inherent binding site of ubiquinone is suggested to overlap with STG-1a that is distinct from the binding site for NADH.
Organizational Affiliation:
Department of Cardiovascular Physiology, Faculty of Medicine, Kagawa University, Kita-gun, Kagawa, 761-0793, Japan. tyama@med.kagawa-u.ac.jp.