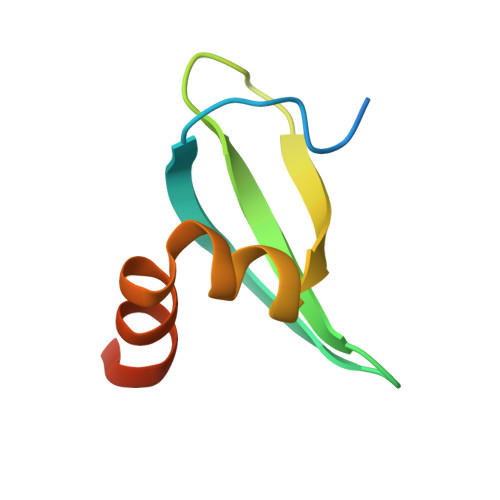
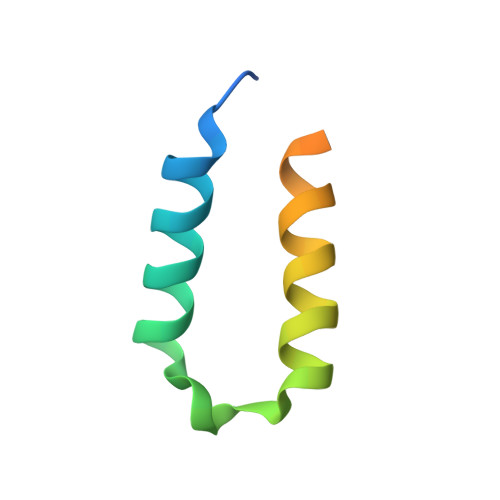
Structural insights into Rhino-Deadlock complex for germline piRNA cluster specification
Yu, B., Lin, Y.A., Parhad, S.S., Jin, Z., Ma, J., Theurkauf, W.E., Zhang, Z.Z., Huang, Y.(2018) EMBO Rep 19
- PubMed: 29858487
- DOI: https://doi.org/10.15252/embr.201745418
- Primary Citation of Related Structures:
5XYV, 5XYW - PubMed Abstract:
PIWI-interacting RNAs (piRNAs) silence transposons in germ cells to maintain genome stability and animal fertility. Rhino, a rapidly evolving heterochromatin protein 1 (HP1) family protein, binds Deadlock in a species-specific manner and so defines the piRNA-producing loci in the Drosophila genome. Here, we determine the crystal structures of Rhino-Deadlock complex in Drosophila melanogaster and simulans In both species, one Rhino binds the N-terminal helix-hairpin-helix motif of one Deadlock protein through a novel interface formed by the beta-sheet in the Rhino chromoshadow domain. Disrupting the interface leads to infertility and transposon hyperactivation in flies. Our structural and functional experiments indicate that electrostatic repulsion at the interaction interface causes cross-species incompatibility between the sibling species. By determining the molecular architecture of this piRNA-producing machinery, we discover a novel HP1-partner interacting mode that is crucial to piRNA biogenesis and transposon silencing. We thus explain the cross-species incompatibility of two sibling species at the molecular level.
Organizational Affiliation:
State Key Laboratory of Molecular Biology, National Center for Protein Science Shanghai, Shanghai Science Research Center, Shanghai Key Laboratory of Molecular Andrology, CAS Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, University of Chinese Academy of Sciences, Shanghai, China.