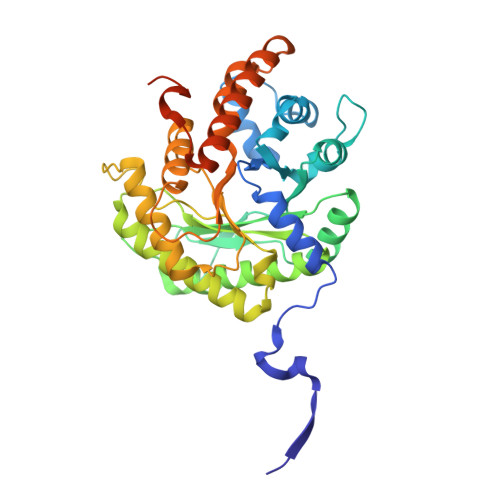
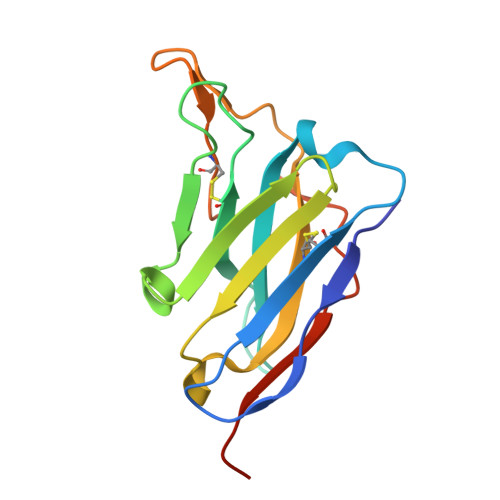
Structural basis for the high specificity of a Trypanosoma congolense immunoassay targeting glycosomal aldolase.
Pinto, J., Odongo, S., Lee, F., Gaspariunaite, V., Muyldermans, S., Magez, S., Sterckx, Y.G.(2017) PLoS Negl Trop Dis 11: e0005932-e0005932
- PubMed: 28915239
- DOI: https://doi.org/10.1371/journal.pntd.0005932
- Primary Citation of Related Structures:
5O0W - PubMed Abstract:
Animal African trypanosomosis (AAT) is a neglected tropical disease which imposes a heavy burden on the livestock industry in Sub-Saharan Africa. Its causative agents are Trypanosoma parasites, with T. congolense and T. vivax being responsible for the majority of the cases. Recently, we identified a Nanobody (Nb474) that was employed to develop a homologous sandwich ELISA targeting T. congolense fructose-1,6-bisphosphate aldolase (TcoALD). Despite the high sequence identity between trypanosomatid aldolases, the Nb474-based immunoassay is highly specific for T. congolense detection. The results presented in this paper yield insights into the molecular principles underlying the assay's high specificity. The structure of the Nb474-TcoALD complex was determined via X-ray crystallography. Together with analytical gel filtration, the structure reveals that a single TcoALD tetramer contains four binding sites for Nb474. Through a comparison with the crystal structures of two other trypanosomatid aldolases, TcoALD residues Ala77 and Leu106 were identified as hot spots for specificity. Via ELISA and surface plasmon resonance (SPR), we demonstrate that mutation of these residues does not abolish TcoALD recognition by Nb474, but does lead to a lack of detection in the Nb474-based homologous sandwich immunoassay. The results show that the high specificity of the Nb474-based immunoassay is not determined by the initial recognition event between Nb474 and TcoALD, but rather by its homologous sandwich design. This (i) provides insights into the optimal set-up of the assay, (ii) may be of great significance for field applications as it could explain the potential detection escape of certain T. congolense strains, and (iii) may be of general interest to those developing similar assays.
Organizational Affiliation:
Research Unit for Cellular and Molecular Immunology (CMIM), Vrije Universiteit Brussel (VUB), Brussels, Belgium.