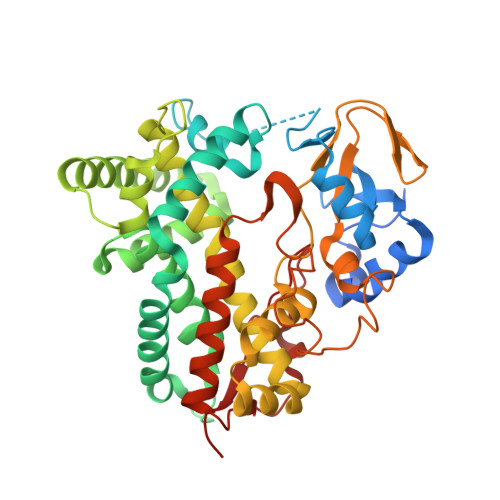
Ketonization of Proline Residues in the Peptide Chains of Actinomycins by a 4-Oxoproline Synthase.
Semsary, S., Crnovcic, I., Driller, R., Vater, J., Loll, B., Keller, U.(2018) Chembiochem 19: 706-715
- PubMed: 29327817
- DOI: https://doi.org/10.1002/cbic.201700666
- Primary Citation of Related Structures:
5NWS - PubMed Abstract:
X-type actinomycins (Acms) contain 4-hydroxyproline (Acm X 0 ) or 4-oxoproline (Acm X 2 ) in their β-pentapeptide lactone rings, whereas their α ring contains proline. We demonstrate that these Acms are formed through asymmetric condensation of Acm half molecules (Acm halves) containing proline with 4-hydroxyproline- or 4-oxoproline-containing Acm halves. In turn, we show-using an artificial Acm half analogue (PPL 1) with proline in its peptide chain-their conversion into the 4-hydroxyproline- and 4-oxoproline-containing Acm halves, PPL 0 and PPL 2, in mycelial suspensions of Streptomyces antibioticus. Two responsible genes of the Acm X biosynthetic gene cluster of S. antibioticus, saacmM and saacmN, encoding a cytochrome P450 monooxygenase (Cyp) and a ferredoxin were identified. After coexpression in Escherichia coli, their gene products converted PPL 1 into PPL 0 and PPL 2 in vivo as well as in situ in permeabilized cell of the transformed E. coli strain in conjunction with the host-encoded ferredoxin reductase in a NADH (NADPH)-dependent manner. saAcmM has high sequence similarity to the Cyp107Z (Ema) family of Cyps, which can convert avermectin B1 into its keto derivative, 4''-oxoavermectin B1. Determination of the structure of saAcmM reveals high similarity to the Ema structure but with significant differences in residues decorating their active sites, which defines saAcmM and its orthologues as a distinct new family of peptidylprolineketonizing Cyp.
Organizational Affiliation:
Institut für Chemie, Biologische Chemie, Technische Universität Berlin, Müller-Breslau-Strasse 10, 10623, Berlin, Germany.